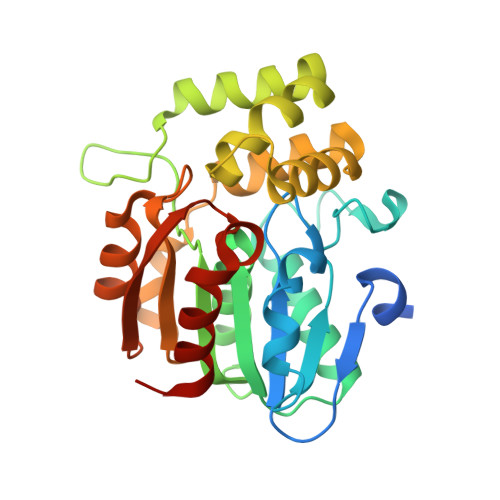
The Structure of the Karrikin-Insensitive Protein (KAI2) in Arabidopsis thaliana
Bythell-Douglas, R., Waters, M.T., Scaffidi, A., Flematti, G.R., Smith, S.M., Bond, C.S.(2013) PLoS One 8: e54758-e54758
- PubMed: 23349965 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0054758
- Primary Citation Related Structures:
4HRX, 4HRY, 4HTA - PubMed Abstract:
KARRIKIN INSENSITIVE 2 (KAI2) is an α/β hydrolase involved in seed germination and seedling development. It is essential for plant responses to karrikins, a class of butenolide compounds derived from burnt plant material that are structurally similar to strigolactone plant hormones. The mechanistic basis for the function of KAI2 in plant development remains unclear. We have determined the crystal structure of Arabidopsis thaliana KAI2 in space groups P2(1) 2(1) 2(1) (a =63.57 Å, b =66.26 Å, c =78.25 Å) and P2(1) (a =50.20 Å, b =56.04 Å, c =52.43 Å, β =116.12°) to 1.55 and 2.11 Å respectively. The catalytic residues are positioned within a large hydrophobic pocket similar to that of DAD2, a protein required for strigolactone response in Petunia hybrida. KAI2 possesses a second solvent-accessible pocket, adjacent to the active site cavity, which offers the possibility of allosteric regulation. The structure of KAI2 is consistent with its designation as a serine hydrolase, as well as previous data implicating the protein in karrikin and strigolactone signalling.
- School of Chemistry and Biochemistry, The University of Western Australia, Crawley, WA, Australia.
Organizational Affiliation: