Molecular mechanism of quinone signaling mediated through S-quinonization of a YodB family repressor QsrR.
Ji, Q., Zhang, L., Jones, M.B., Sun, F., Deng, X., Liang, H., Cho, H., Brugarolas, P., Gao, Y.N., Peterson, S.N., Lan, L., Bae, T., He, C.(2013) Proc Natl Acad Sci U S A 110: 5010-5015
- PubMed: 23479646 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1219446110
- Primary Citation Related Structures:
4HQE, 4HQM - PubMed Abstract:
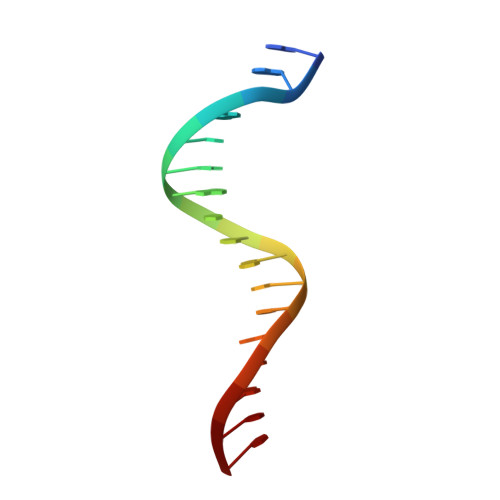
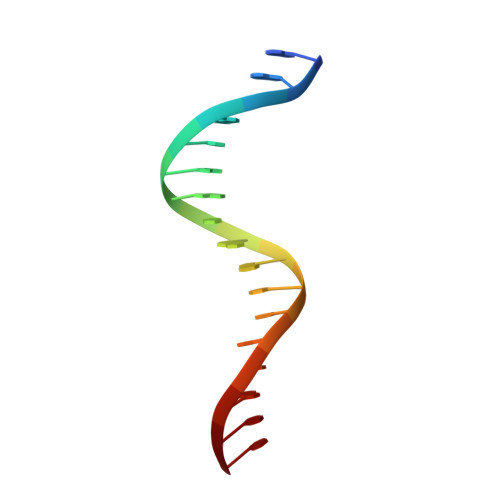
Quinone molecules are intracellular electron-transport carriers, as well as critical intra- and extracellular signals. However, transcriptional regulation of quinone signaling and its molecular basis are poorly understood. Here, we identify a thiol-stress-sensing regulator YodB family transcriptional regulator as a central component of quinone stress response of Staphylococcus aureus, which we have termed the quinone-sensing and response repressor (QsrR). We also identify and confirm an unprecedented quinone-sensing mechanism based on the S-quinonization of the essential residue Cys-5. Structural characterizations of the QsrR-DNA and QsrR-menadione complexes further reveal that the covalent association of menadione directly leads to the release of QsrR from operator DNA following a 10° rigid-body rotation as well as a 9-Å elongation between the dimeric subunits. The molecular level characterization of this quinone-sensing transcriptional regulator provides critical insights into quinone-mediated gene regulation in human pathogens.
- Department of Chemistry, The University of Chicago, Chicago, IL 60637, USA.
Organizational Affiliation: