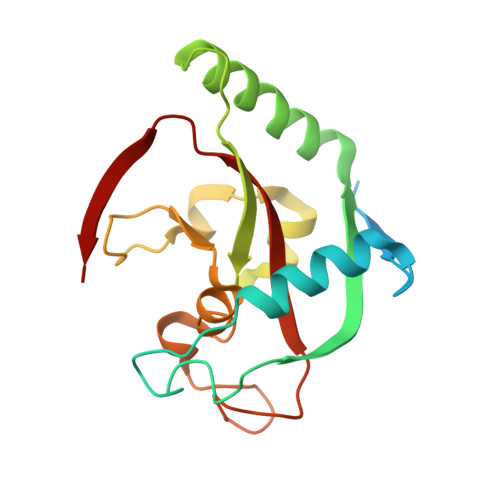
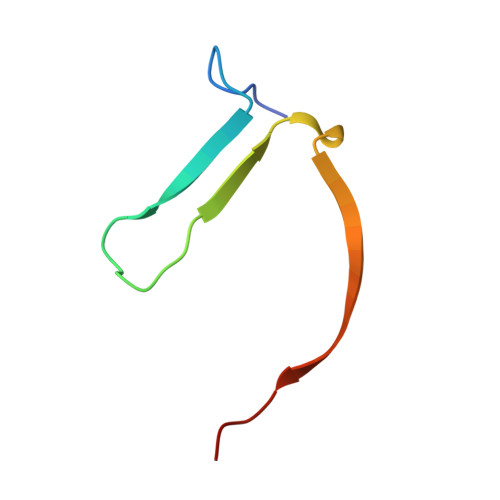
Screening and structural analysis of flavones inhibiting tankyrases.
Narwal, M., Haikarainen, T., Fallarero, A., Vuorela, P.M., Lehtio, L.(2013) J Med Chem 56: 3507-3517
- PubMed: 23574272 Search on PubMed
- DOI: https://doi.org/10.1021/jm3018783
- Primary Citation Related Structures:
4HKI, 4HKK, 4HKN, 4HL5, 4HLF, 4HLG, 4HLH, 4HLK, 4HLM, 4HMH - PubMed Abstract:
Flavonoids are known for their beneficial effects on human health, and therefore the therapeutic potential of these compounds have been extensively studied. Flavone has been previously identified as a tankyrase inhibitor, and to further elucidate whether tankyrases would be inhibited by other flavonoids, we performed a systematic screening of tankyrase 2 inhibitory activity using 500 natural and naturally derived flavonoids covering nine different flavonoid classes. All identified tankyrase inhibitors were flavones. We report crystal structures of all the hit compounds in complex with the catalytic domain of human tankyrase 2. Flavone derivatives in all 10 crystal structures bind to the nicotinamide binding site of tankyrase 2. Potencies of the active flavones toward tankyrases vary between 50 nM and 1.1 μM, and flavones show up to 200-fold selectivity for tankyrases over ARTD1. The molecular details of the interactions revealed by cocrystal structures efficiently describe the properties of potent flavone derivatives inhibiting tankyrases.
- Biocenter Oulu and Department of Biochemistry, University of Oulu, Oulu, Finland.
Organizational Affiliation: