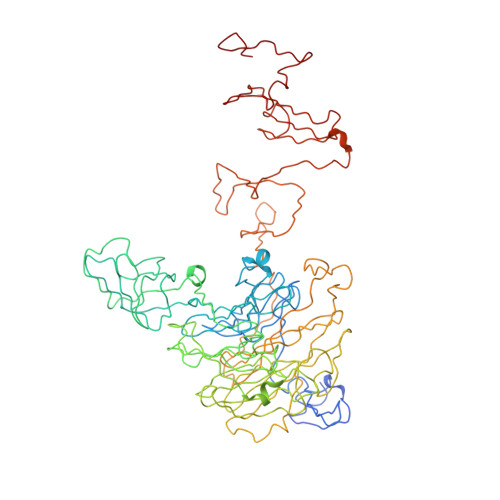
Structure and biochemical characterization of bacteriophage phi92 endosialidase.
Schwarzer, D., Browning, C., Stummeyer, K., Oberbeck, A., Muhlenhoff, M., Gerardy-Schahn, R., Leiman, P.G.(2015) Virology 477: 133-143
- PubMed: 25475852 Search on PubMed
- DOI: https://doi.org/10.1016/j.virol.2014.11.002
- Primary Citation Related Structures:
4HIZ - PubMed Abstract:
Surface-associated capsular polysaccharides (CPSs) protect bacteria against phage infection and enhance pathogenicity by interfering with the function of the host innate immune system. The CPS of enteropathogenic Escherichia coli K92 is a unique sialic acid polymer (polySia) with alternating α2,8- and α2,9-linkages. This CPS can be digested by the gene 143 encoded endosialidase of bacteriophage phi92. Here we report the crystal structure of the phi92 endosialidase in complex with a dimer of α2,9-linked sialic acid and analyze its catalytic functions. Unlike the well characterized and homologous endosialidase of phage K1F, the phi92 endosialidase is a bifunctional enzyme with high activity against α2,8- and low activity against α2,9-linkages in a polySia chain. Moreover, in contrast to the processive K1F endosialidase, the phi92 endosialidase degrades the polymer in a non-processive mode. Beyond describing the first endosialidase with α2,9-specificity, our data introduce a novel platform for studies of endosialidase regioselectivity and for engineering highly active α2,9-specific enzymes.
- Institute for Cellular Chemistry, Hannover Medical School (MHH), 30625 Hannover, Germany.
Organizational Affiliation: