Structure of integrin-linked kinase ankyrin repeat domain in complex with PINCH1 LIM1 domain collected at high energy, wavelength 0.32800
Stiegler, A.L., Jakoncic, J., Stojanoff, V., Chiswell, B.P., Calderwood, D.A., Boggon, T.J.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Integrin-linked protein kinase | 179 | Homo sapiens | Mutation(s): 0 Gene Names: ILK, ILK1, ILK2 EC: 2.7.11.1 | 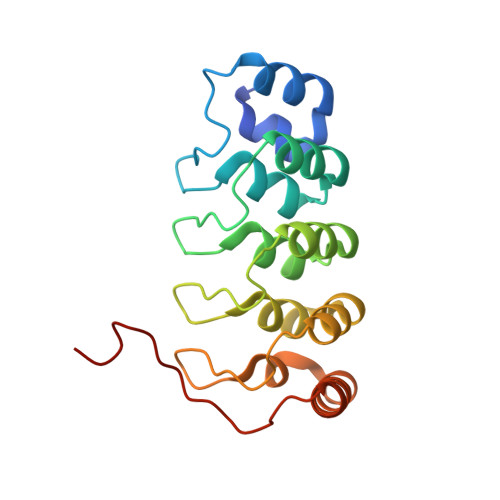 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q13418 GTEx: ENSG00000166333 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q13418 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| LIM and senescent cell antigen-like-containing domain protein 1 | 72 | Homo sapiens | Mutation(s): 0 Gene Names: LIMS1, PINCH, PINCH1 | 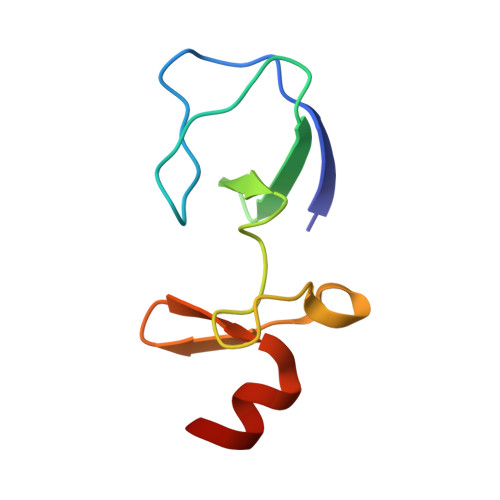 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P48059 GTEx: ENSG00000169756 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P48059 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PO4 Download:Ideal Coordinates CCD File | C [auth A] | PHOSPHATE ION O4 P NBIIXXVUZAFLBC-UHFFFAOYSA-K |  | ||
| ZN Download:Ideal Coordinates CCD File | D [auth B], E [auth B] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 37.093 | α = 79.87 |
| b = 41.859 | β = 68.82 |
| c = 46.983 | γ = 85.77 |
| Software Name | Purpose |
|---|---|
| DENZO | data reduction |
| SCALEPACK | data scaling |
| PHASER | phasing |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |