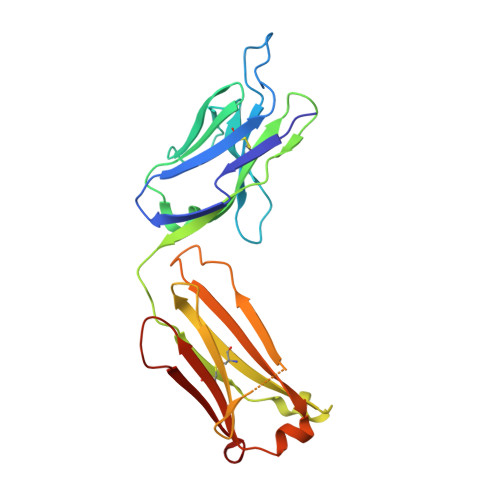
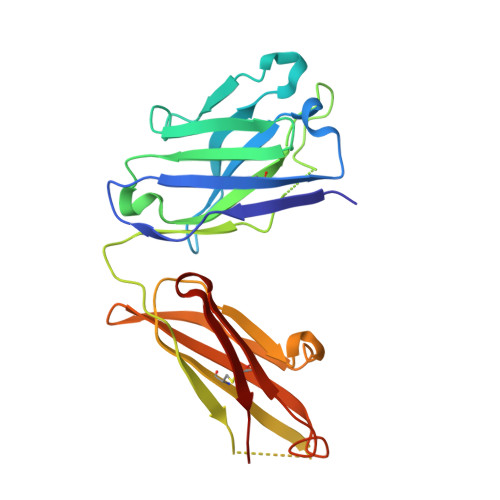
Exploring the cross-reactivity of S25-2: complex with a 5,6-dehydro-Kdo disaccharide.
Brooks, C.L., Wimmer, K., Kosma, P., Muller-Loennies, S., Brade, L., Brade, H., Evans, S.V.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 2-5
- PubMed: 23295476 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309112047422
- Primary Citation Related Structures:
4HGW - PubMed Abstract:
The near-germline antibody S25-2 exhibits a remarkable cross-reactivity for oligosaccharides containing the bacterial lipopolysaccharide carbohydrate 3-deoxy-D-manno-oct-2-ulosonic acid (Kdo). The recent synthesis of a variety of Kdo analogues permits a detailed structural analysis of the importance of specific interactions in antigen recognition by S25-2. The Kdo disaccharide analogue Kdo-(2→4)-5,6-dehydro-Kdo lacks a 5-OH group on the second Kdo residue and has been cocrystallized with S25-2. The structure reveals that the modification of the Kdo residue at position 5 results in a rearrangement of intramolecular hydrogen bonds in the antigen that allows it to assume a novel conformation in the antibody-combining site. The cross-reactive binding of S25-2 to this synthetic ligand highlights the adaptability of this antibody to non-natural synthetic analogues.
- Department of Biochemistry and Microbiology, University of Victoria, PO Box 3055 STN CSC, Victoria, BC V8W 3P6, Canada. cbrooks1@ualberta.ca
Organizational Affiliation: