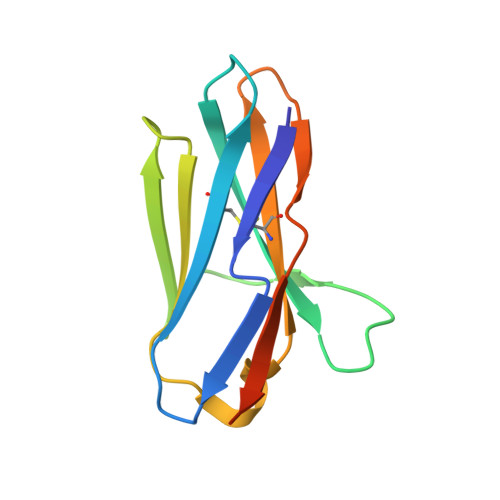
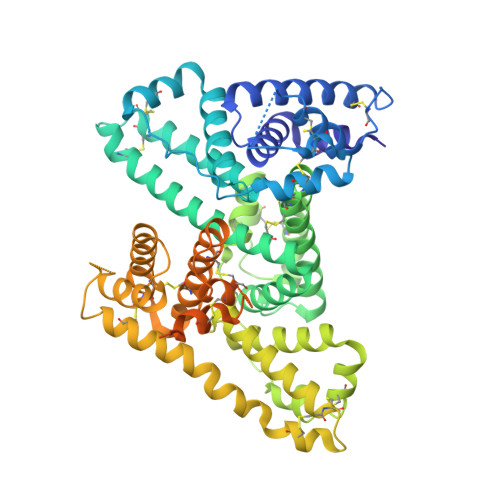
Atypical Antigen Recognition Mode of a Shark Immunoglobulin New Antigen Receptor (IgNAR) Variable Domain Characterized by Humanization and Structural Analysis.
Kovalenko, O.V., Olland, A., Piche-Nicholas, N., Godbole, A., King, D., Svenson, K., Calabro, V., Muller, M.R., Barelle, C.J., Somers, W., Gill, D.S., Mosyak, L., Tchistiakova, L.(2013) J Biological Chem 288: 17408-17419
- PubMed: 23632026 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.435289
- Primary Citation Related Structures:
4HGK, 4HGM - PubMed Abstract:
The immunoglobulin new antigen receptors (IgNARs) are a class of Ig-like molecules of the shark immune system that exist as heavy chain-only homodimers and bind antigens by their single domain variable regions (V-NARs). Following shark immunization and/or in vitro selection, V-NARs can be generated as soluble, stable, and specific high affinity monomeric binding proteins of ∼12 kDa. We have previously isolated a V-NAR from an immunized spiny dogfish shark, named E06, that binds specifically and with high affinity to human, mouse, and rat serum albumins. Humanization of E06 was carried out by converting over 60% of non-complementarity-determining region residues to those of a human germ line Vκ1 sequence, DPK9. The resulting huE06 molecules have largely retained the specificity and affinity of antigen binding of the parental V-NAR. Crystal structures of the shark E06 and its humanized variant (huE06 v1.1) in complex with human serum albumin (HSA) were determined at 3- and 2.3-Å resolution, respectively. The huE06 v1.1 molecule retained all but one amino acid residues involved in the binding site for HSA. Structural analysis of these V-NARs has revealed an unusual variable domain-antigen interaction. E06 interacts with HSA in an atypical mode that utilizes extensive framework contacts in addition to complementarity-determining regions that has not been seen previously in V-NARs. On the basis of the structure, the roles of various elements of the molecule are described with respect to antigen binding and V-NAR stability. This information broadens the general understanding of antigen recognition and provides a framework for further design and humanization of shark IgNARs.
- Global Biotherapeutics Technologies, Pfizer Research and Development, Cambridge, Massachusetts 02140, USA. oleg.kovalenko@pfizer.com
Organizational Affiliation: