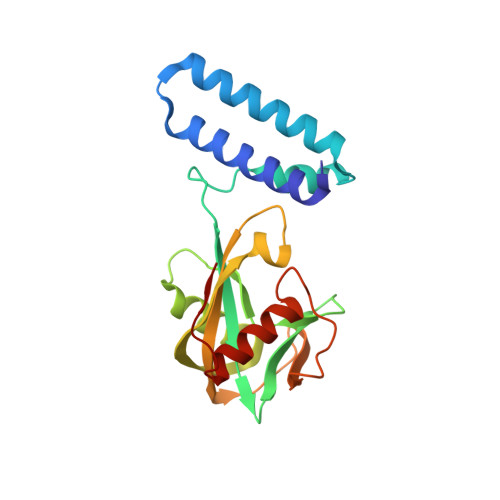
A UDP-X diphosphatase from Streptococcus pneumoniae hydrolyzes precursors of peptidoglycan biosynthesis.
Duong-Ly, K.C., Woo, H.N., Dunn, C.A., Xu, W., Babic, A., Bessman, M.J., Amzel, L.M., Gabelli, S.B.(2013) PLoS One 8: e64241-e64241
- PubMed: 23691178 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0064241
- Primary Citation Related Structures:
4HFQ - PubMed Abstract:
The gene for a Nudix enzyme (SP_1669) was found to code for a UDP-X diphosphatase. The SP_1669 gene is localized among genes encoding proteins that participate in cell division in Streptococcus pneumoniae. One of these genes, MurF, encodes an enzyme that catalyzes the last step of the Mur pathway of peptidoglycan biosynthesis. Mur pathway substrates are all derived from UDP-glucosamine and all are potential Nudix substrates. We showed that UDP-X diphosphatase can hydrolyze the Mur pathway substrates UDP-N-acetylmuramic acid and UDP-N-acetylmuramoyl-L-alanine. The 1.39 Å resolution crystal structure of this enzyme shows that it folds as an asymmetric homodimer with two distinct active sites, each containing elements of the conserved Nudix box sequence. In addition to its Nudix catalytic activity, the enzyme has a 3'5' RNA exonuclease activity. We propose that the structural asymmetry in UDP-X diphosphatase facilitates the recognition of these two distinct classes of substrates, Nudix substrates and RNA. UDP-X diphosphatase is a prototype of a new family of Nudix enzymes with unique structural characteristics: two monomers, each consisting of an N-terminal helix bundle domain and a C-terminal Nudix domain, form an asymmetric dimer with two distinct active sites. These enzymes function to hydrolyze bacterial cell wall precursors and degrade RNA.
- Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, Baltimore, Maryland, United States of America.
Organizational Affiliation: