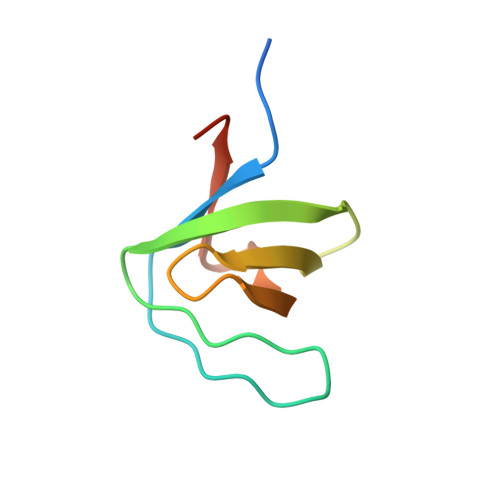
Solution structure of the human Hck SH3 domain and identification of its ligand binding site.
Horita, D.A., Baldisseri, D.M., Zhang, W., Altieri, A.S., Smithgall, T.E., Gmeiner, W.H., Byrd, R.A.(1998) J Mol Biology 278: 253-265
- PubMed: 9571048 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1998.1690
- Primary Citation Related Structures:
4HCK, 5HCK - PubMed Abstract:
SH3 domains are protein binding domains that occur widely among signal transduction proteins. Here, we present the NMR-determined solution structure of the SH3 domain from the cytoplasmic protein tyrosine kinase, Hck. Hck is involved in a number of cell signal transduction pathways, frequently in pathways associated with immune response. SH3 domains bind proteins via a left-handed polyproline type II helix on the target protein. We have assessed the structural impact of binding to a ligand through addition of a peptide corresponding to a proline-rich region of a Hck target, the GTPase activating protein of the Ras pathway. Ligand binding effects small structural changes and stabilizes the SH3 domain structure. Also, we have compared the solution structure of the Hck SH3 domain to the crystal structure of Hck, in which the SH3 domain exhibits an intramolecular binding to an interdomain linker region. These structures are interpreted as the apo- and holo- forms of the Hck SH3 domain.
- ABL-Basic Research Program, NCI-Frederick Cancer Research and Development Center, Frederick, MD, 21702-1201, USA.
Organizational Affiliation: