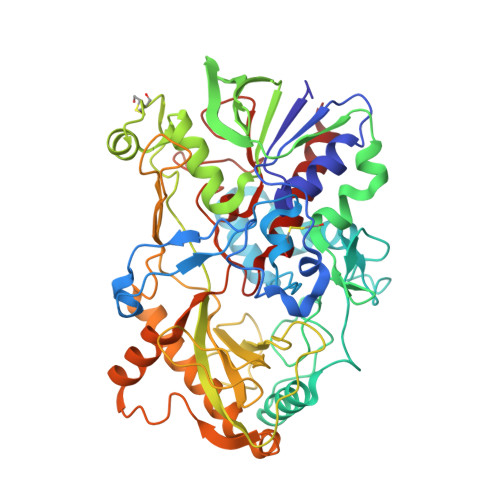
Crystal structure of pyridoxine 4-oxidase from Mesorhizobium loti.
Mugo, A.N., Kobayashi, J., Yamasaki, T., Mikami, B., Ohnishi, K., Yoshikane, Y., Yagi, T.(2013) Biochim Biophys Acta 1834: 953-963
- PubMed: 23501672 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2013.03.004
- Primary Citation Related Structures:
4HA6 - PubMed Abstract:
Pyridoxine 4-oxidase (PNOX) from Mesorhizobium loti is a monomeric glucose-methanol-choline (GMC) oxidoreductase family enzyme, catalyzes FAD-dependent oxidation of pyridoxine (PN) into pyridoxal, and is the first enzyme in pathway I for the degradation of PN. The tertiary structures of PNOX with a C-terminal His6-tag and PNOX-pyridoxamine (PM) complex were determined at 2.2Å and at 2.1Å resolutions, respectively. The overall structure consisted of FAD-binding and substrate-binding domains. In the active site, His460, His462, and Pro504 were located on the re-face of the isoalloxazine ring of FAD. PM binds to the active site through several hydrogen bonds. The side chains of His462 and His460 are located at 2.7 and 3.1Å from the N4' atom of PM. The activities of His460Ala and His462Ala mutant PNOXs were very low, and 460Ala/His462Ala double mutant PNOX exhibited no activity. His462 may act as a general base for the abstraction of a proton from the 4'-hydroxyl of PN. His460 may play a role in the binding and positioning of PN. The C4' atom in PM is located at 3.2Å, and the hydride ion from the C4' atom may be transferred to the N5 atom of the isoalloxazine ring. The comparison of active site residues in GMC oxidoreductase shows that Pro504 in PNOX corresponds to Asn or His of the conserved His-Asn or His-His pair in other GMC oxidoreductases. The function of the novel proline residue was discussed.
- Graduate School of Integral Arts and Science, Kochi University, Nankoku, Kochi, Japan.
Organizational Affiliation: