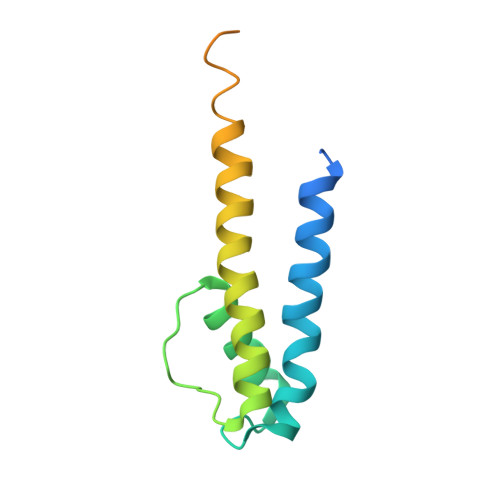
Crystal Structure of a Voltage-gated K+ Channel Pore Module in a Closed State in Lipid Membranes.
Santos, J.S., Asmar-Rovira, G.A., Han, G.W., Liu, W., Syeda, R., Cherezov, V., Baker, K.A., Stevens, R.C., Montal, M.(2012) J Biological Chem 287: 43063-43070
- PubMed: 23095758 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.415091
- Primary Citation Related Structures:
4H33, 4H37 - PubMed Abstract:
Voltage-gated K(+) channels underlie the electrical excitability of cells. Each subunit of the functional tetramer consists of the tandem fusion of two modules, an N-terminal voltage-sensor and a C-terminal pore. To investigate how sensor coupling to the pore generates voltage-dependent channel opening, we solved the crystal structure and characterized the function of a voltage-gated K(+) channel pore in a lipid membrane. The structure of a functional channel in a membrane environment at 3.1 Å resolution establishes an unprecedented connection between channel structure and function. The structure is unique in delineating an ion-occupied ready to conduct selectivity filter, a confined aqueous cavity, and a closed activation gate, embodying a dynamic entity trapped in an unstable closed state.
- Section of Neurobiology, Division of Biological Sciences, University of California San Diego, La Jolla, California 92093, USA.
Organizational Affiliation: