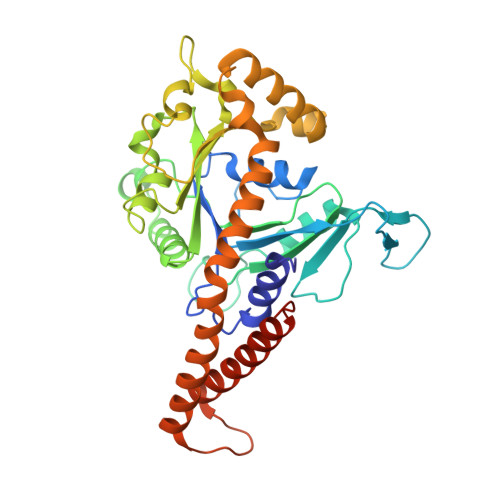
Functional Mapping of Human Dynamin-1-Like GTPase Domain Based on X-ray Structure Analyses.
Wenger, J., Klinglmayr, E., Frohlich, C., Eibl, C., Gimeno, A., Hessenberger, M., Puehringer, S., Daumke, O., Goettig, P.(2013) PLoS One 8: e71835-e71835
- PubMed: 23977156 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0071835
- Primary Citation Related Structures:
4H1U, 4H1V - PubMed Abstract:
Human dynamin-1-like protein (DNM1L) is a GTP-driven molecular machine that segregates mitochondria and peroxisomes. To obtain insights into its catalytic mechanism, we determined crystal structures of a construct comprising the GTPase domain and the bundle signaling element (BSE) in the nucleotide-free and GTP-analogue-bound states. The GTPase domain of DNM1L is structurally related to that of dynamin and binds the nucleotide 5'-Guanylyl-imidodiphosphate (GMP-PNP) via five highly conserved motifs, whereas the BSE folds into a pocket at the opposite side. Based on these structures, the GTPase center was systematically mapped by alanine mutagenesis and kinetic measurements. Thus, residues essential for the GTPase reaction were characterized, among them Lys38, Ser39 and Ser40 in the phosphate binding loop, Thr59 from switch I, Asp146 and Gly149 from switch II, Lys216 and Asp218 in the G4 element, as well as Asn246 in the G5 element. Also, mutated Glu81 and Glu82 in the unique 16-residue insertion of DNM1L influence the activity significantly. Mutations of Gln34, Ser35, and Asp190 in the predicted assembly interface interfered with dimerization of the GTPase domain induced by a transition state analogue and led to a loss of the lipid-stimulated GTPase activity. Our data point to related catalytic mechanisms of DNM1L and dynamin involving dimerization of their GTPase domains.
- Department of Molecular Biology, University of Salzburg, Salzburg, Austria.
Organizational Affiliation: