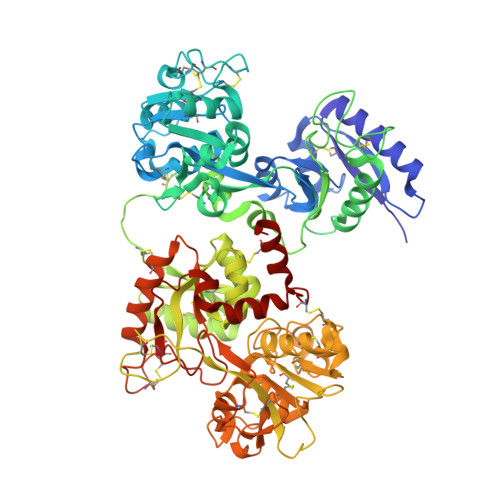
Iron and bismuth bound human serum transferrin reveals a partially-opened conformation in the N-lobe.
Yang, N., Zhang, H., Wang, M., Hao, Q., Sun, H.(2012) Sci Rep 2: 999-999
- PubMed: 23256035
- DOI: https://doi.org/10.1038/srep00999
- Primary Citation of Related Structures:
3QYT, 4H0W - PubMed Abstract:
Human serum transferrin (hTF) binds Fe(III) tightly but reversibly, and delivers it to cells via a receptor-mediated endocytosis process. The metal-binding and release result in significant conformational changes of the protein. Here, we report the crystal structures of diferric-hTF (Fe(N)Fe(C)-hTF) and bismuth-bound hTF (Bi(N)Fe(C)-hTF) at 2.8 and 2.4 Å resolutions respectively. Notably, the N-lobes of both structures exhibit unique "partially-opened" conformations between those of the apo-hTF and holo-hTF. Fe(III) and Bi(III) in the N-lobe coordinate to, besides anions, only two (Tyr95 and Tyr188) and one (Tyr188) tyrosine residues, respectively, in contrast to four residues in the holo-hTF. The C-lobe of both structures are fully closed with iron coordinating to four residues and a carbonate. The structures of hTF observed here represent key conformers captured in the dynamic nature of the transferrin family proteins and provide a structural basis for understanding the mechanism of metal uptake and release in transferrin families.
- Department of Chemistry, the University of Hong Kong, Pokfulam Road, Hong Kong, PR China.
Organizational Affiliation: