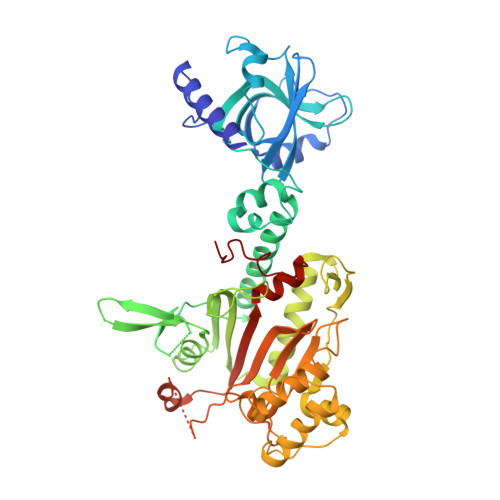
Structural analysis of malaria-parasite lysyl-tRNA synthetase provides a platform for drug development.
Khan, S., Garg, A., Camacho, N., Van Rooyen, J., Kumar Pole, A., Belrhali, H., Ribas de Pouplana, L., Sharma, V., Sharma, A.(2013) Acta Crystallogr D Biol Crystallogr 69: 785-795
- PubMed: 23633587 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444913001923
- Primary Citation Related Structures:
4H02 - PubMed Abstract:
Aminoacyl-tRNA synthetases are essential enzymes that transmit information from the genetic code to proteins in cells and are targets for antipathogen drug development. Elucidation of the crystal structure of cytoplasmic lysyl-tRNA synthetase from the malaria parasite Plasmodium falciparum (PfLysRS) has allowed direct comparison with human LysRS. The authors' data suggest that PfLysRS is dimeric in solution, whereas the human counterpart can also adopt tetrameric forms. It is shown for the first time that PfLysRS is capable of synthesizing the signalling molecule Ap4a (diadenosine tetraphosphate) using ATP as a substrate. The PfLysRS crystal structure is in the apo form, such that binding to ATP will require rotameric changes in four conserved residues. Differences in the active-site regions of parasite and human LysRSs suggest the possibility of exploiting PfLysRS for selective inhibition. These investigations on PfLysRS further validate malarial LysRSs as attractive antimalarial targets and provide new structural space for the development of inhibitors that target pathogen LysRSs selectively.
- Structural and Computational Biology Group, International Centre for Genetic Engineering and Biotechnology, Aruna Asaf Ali Road, New Delhi 110 067, India.
Organizational Affiliation: