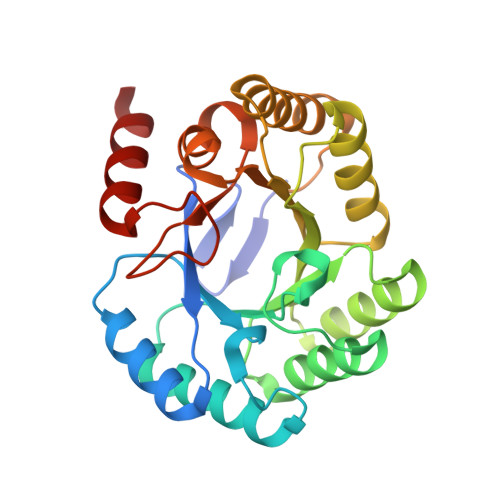
Crystal structures of type I dehydroquinate dehydratase in complex with quinate and shikimate suggest a novel mechanism of schiff base formation.
Light, S.H., Antanasijevic, A., Krishna, S.N., Caffrey, M., Anderson, W.F., Lavie, A.(2014) Biochemistry 53: 872-880
- PubMed: 24437575 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi4015506
- Primary Citation Related Structures:
4GUI, 4GUJ, 4IUO - PubMed Abstract:
A component of the shikimate biosynthetic pathway, dehydroquinate dehydratase (DHQD) catalyzes the dehydration of 3-dehydroquniate (DHQ) to 3-dehydroshikimate. In the type I DHQD reaction mechanism a lysine forms a Schiff base intermediate with DHQ. The Schiff base acts as an electron sink to facilitate the catalytic dehydration. To address the mechanism of Schiff base formation, we determined structures of the Salmonella enterica wild-type DHQD in complex with the substrate analogue quinate and the product analogue shikimate. In addition, we determined the structure of the K170M mutant (Lys170 being the Schiff base forming residue) in complex with quinate. Combined with nuclear magnetic resonance and isothermal titration calorimetry data that revealed altered binding of the analogue to the K170M mutant, these structures suggest a model of Schiff base formation characterized by the dynamic interplay of opposing forces acting on either side of the substrate. On the side distant from the substrate 3-carbonyl group, closure of the enzyme's β8-α8 loop is proposed to guide DHQ into the proximity of the Schiff base-forming Lys170. On the 3-carbonyl side of the substrate, Lys170 sterically alters the position of DHQ's reactive ketone, aligning it at an angle conducive for nucleophilic attack. This study of a type I DHQD reveals the interplay between the enzyme and substrate required for the correct orientation of a functional group constrained within a cyclic substrate.
- Center for Structural Genomics of Infectious Diseases and Department of Molecular Pharmacology and Biological Chemistry, Feinberg School of Medicine, Northwestern University , Chicago, Illinois 60611, United States.
Organizational Affiliation: