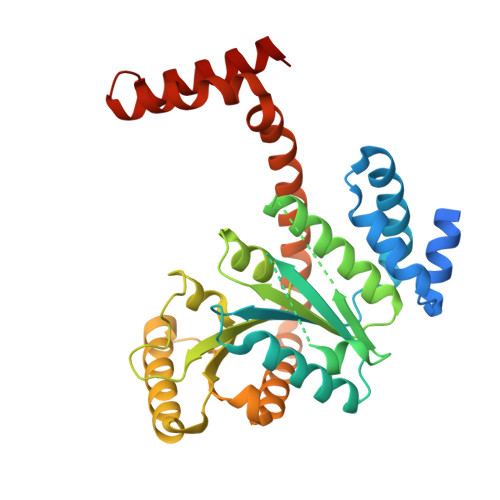
Crystal structures of Mycobacterial MeaB and MMAA-like GTPases.
Edwards, T.E., Baugh, L., Bullen, J., Baydo, R.O., Witte, P., Thompkins, K., Phan, I.Q., Abendroth, J., Clifton, M.C., Sankaran, B., Van Voorhis, W.C., Myler, P.J., Staker, B.L., Grundner, C., Lorimer, D.D.(2015) J Struct Funct Genomics 16: 91-99
- PubMed: 25832174 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s10969-015-9197-2
- Primary Citation Related Structures:
3MD0, 3NXS, 3P32, 3TK1, 4GT1 - PubMed Abstract:
The methylmalonyl Co-A mutase-associated GTPase MeaB from Methylobacterium extorquens is involved in glyoxylate regulation and required for growth. In humans, mutations in the homolog methylmalonic aciduria associated protein (MMAA) cause methylmalonic aciduria, which is often fatal. The central role of MeaB from bacteria to humans suggests that MeaB is also important in other, pathogenic bacteria such as Mycobacterium tuberculosis. However, the identity of the mycobacterial MeaB homolog is presently unclear. Here, we identify the M. tuberculosis protein Rv1496 and its homologs in M. smegmatis and M. thermoresistibile as MeaB. The crystal structures of all three homologs are highly similar to MeaB and MMAA structures and reveal a characteristic three-domain homodimer with GDP bound in the G domain active site. A structure of Rv1496 obtained from a crystal grown in the presence of GTP exhibited electron density for GDP, suggesting GTPase activity. These structures identify the mycobacterial MeaB and provide a structural framework for therapeutic targeting of M. tuberculosis MeaB.
- Beryllium, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Bainbridge Island, WA, 98110, USA, tedwards@be4.com.
Organizational Affiliation: