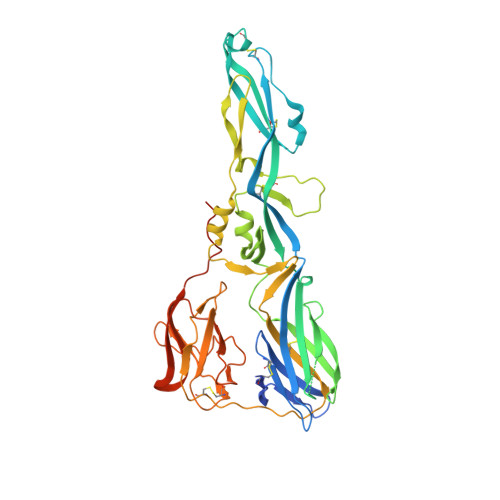
Structure of a dengue virus envelope protein late-stage fusion intermediate.
Klein, D.E., Choi, J.L., Harrison, S.C.(2013) J Virol 87: 2287-2293
- PubMed: 23236058 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.02957-12
- Primary Citation Related Structures:
4GSX, 4GT0 - PubMed Abstract:
The final stages of dengue virus fusion are thought to occur when the membrane-proximal stem drives the transmembrane anchor of the viral envelope protein (E) toward the fusion loop, buried in the target cell membrane. Crystal structures of E have lacked this essential stem region. We expressed and crystallized soluble mutant forms of the dengue virus envelope protein (sE) that include portions of the juxtamembrane stem. Their structures represent late-stage fusion intermediates. The proximal part of the stem has both intra- and intermolecular interactions, so the chain "zips up" along the trimer seam. The penultimate interaction we detected involves the conserved residue F402, which has hydrophobic contacts with a conserved surface on domain II. These interactions do not require any larger-scale changes in trimer packing. The techniques for expression and crystallization of sE containing stem reported here may allow further characterization of the final stages of flavivirus fusion.
- Jack and Eileen Connors Structural Biology Laboratory, Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, Massachusetts, USA.
Organizational Affiliation: