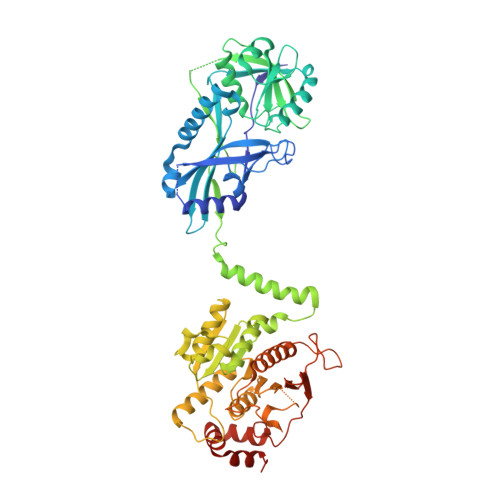
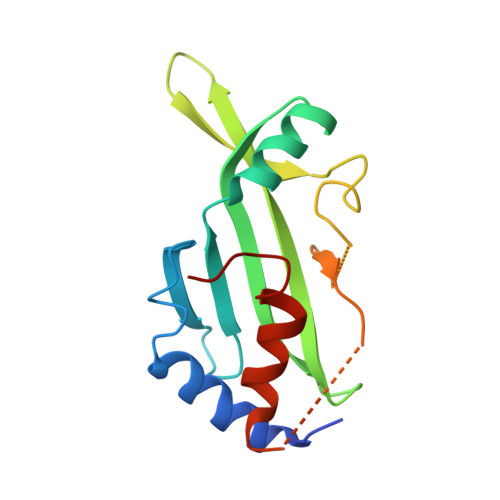
Noncanonical E2 recruitment by the autophagy E1 revealed by Atg7-Atg3 and Atg7-Atg10 structures.
Kaiser, S.E., Mao, K., Taherbhoy, A.M., Yu, S., Olszewski, J.L., Duda, D.M., Kurinov, I., Deng, A., Fenn, T.D., Klionsky, D.J., Schulman, B.A.(2012) Nat Struct Mol Biol 19: 1242-1249
- PubMed: 23142976 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2415
- Primary Citation Related Structures:
4GSJ, 4GSK, 4GSL - PubMed Abstract:
Core functions of autophagy are mediated by ubiquitin-like protein (UBL) cascades, in which a homodimeric E1 enzyme, Atg7, directs the UBLs Atg8 and Atg12 to their respective E2 enzymes, Atg3 and Atg10. Crystallographic and mutational analyses of yeast (Atg7-Atg3)(2) and (Atg7-Atg10)(2) complexes reveal noncanonical, multisite E1-E2 recognition in autophagy. Atg7's unique N-terminal domain recruits distinctive elements from the Atg3 and Atg10 'backsides'. This, along with E1 and E2 conformational variability, allows presentation of 'frontside' Atg3 and Atg10 active sites to the catalytic cysteine in the C-terminal domain from the opposite Atg7 protomer in the homodimer. Despite different modes of binding, the data suggest that common principles underlie conjugation in both noncanonical and canonical UBL cascades, whereby flexibly tethered E1 domains recruit E2s through surfaces remote from their active sites to juxtapose the E1 and E2 catalytic cysteines.
- Department of Structural Biology, St. Jude Children's Research Hospital, Memphis, Tennessee, USA.
Organizational Affiliation: