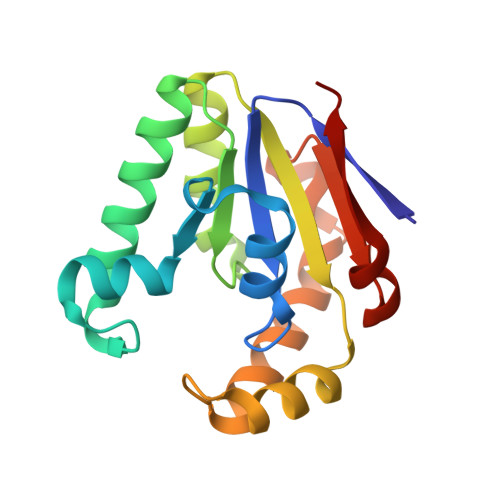
Structure and mechanism of the polynucleotide kinase component of the bacterial Pnkp-Hen1 RNA repair system.
Wang, L.K., Das, U., Smith, P., Shuman, S.(2012) RNA 18: 2277-2286
- PubMed: 23118415 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.036061.112
- Primary Citation Related Structures:
4GP6, 4GP7 - PubMed Abstract:
Pnkp is the end-healing and end-sealing component of an RNA repair system present in diverse bacteria from many phyla. Pnkp is composed of three catalytic modules: an N-terminal polynucleotide 5'-kinase, a central 2',3' phosphatase, and a C-terminal ligase. Here we report the crystal structure of the kinase domain of Clostridium thermocellum Pnkp bound to ATP•Mg²⁺ (substrate complex) and ADP•Mg²⁺ (product complex). The protein consists of a core P-loop phosphotransferase fold embellished by a distinctive homodimerization module composed of secondary structure elements derived from the N and C termini of the kinase domain. ATP is bound within a crescent-shaped groove formed by the P-loop (¹⁵GSSGSGKST²³) and an overlying helix-loop-helix "lid." The α and β phosphates are engaged by a network of hydrogen bonds from Thr23 and the P-loop main-chain amides; the γ phosphate is anchored by the lid residues Arg120 and Arg123. The P-loop lysine (Lys21) and the catalytic Mg²⁺ bridge the ATP β and γ phosphates. The P-loop serine (Ser22) is the sole enzymic constituent of the octahedral metal coordination complex. Structure-guided mutational analysis underscored the essential contributions of Lys21 and Ser22 in the ATP donor site and Asp38 and Arg41 in the phosphoacceptor site. Our studies suggest a catalytic mechanism whereby Asp38 (as general base) activates the polynucleotide 5'-OH for its nucleophilic attack on the γ phosphorus and Lys21 and Mg²⁺ stabilize the transition state.
- Molecular Biology Program, Sloan-Kettering Institute, New York, New York 10065, USA.
Organizational Affiliation: