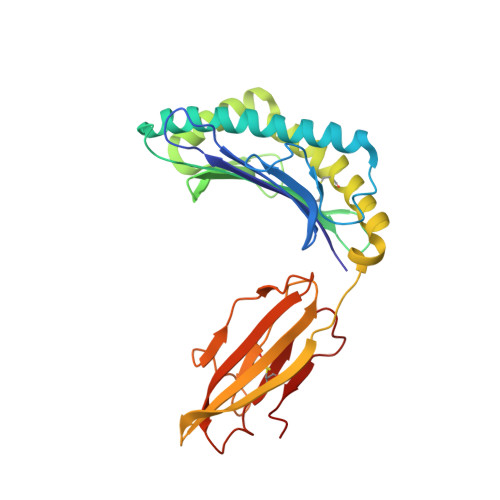
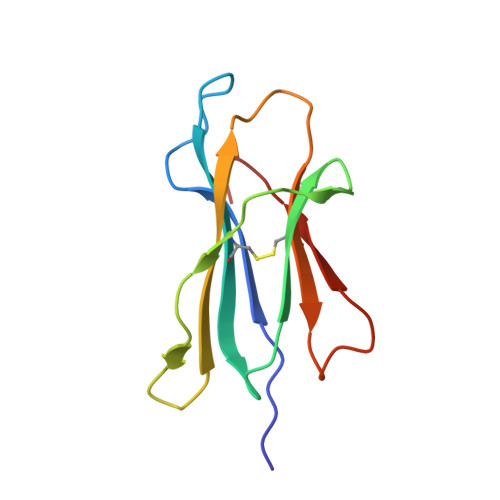
T-cell Receptor-optimized Peptide Skewing of the T-cell Repertoire Can Enhance Antigen Targeting.
Ekeruche-Makinde, J., Clement, M., Cole, D.K., Edwards, E.S., Ladell, K., Miles, J.J., Matthews, K.K., Fuller, A., Lloyd, K.A., Madura, F., Dolton, G.M., Pentier, J., Lissina, A., Gostick, E., Baxter, T.K., Baker, B.M., Rizkallah, P.J., Price, D.A., Wooldridge, L., Sewell, A.K.(2012) J Biological Chem 287: 37269-37281
- PubMed: 22952231 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.386409
- Primary Citation Related Structures:
4GKN, 4GKS - PubMed Abstract:
Altered peptide antigens that enhance T-cell immunogenicity have been used to improve peptide-based vaccination for a range of diseases. Although this strategy can prime T-cell responses of greater magnitude, the efficacy of constituent T-cell clonotypes within the primed population can be poor. To overcome this limitation, we isolated a CD8(+) T-cell clone (MEL5) with an enhanced ability to recognize the HLA A*0201-Melan A(27-35) (HLA A*0201-AAGIGILTV) antigen expressed on the surface of malignant melanoma cells. We used combinatorial peptide library screening to design an optimal peptide sequence that enhanced functional activation of the MEL5 clone, but not other CD8(+) T-cell clones that recognized HLA A*0201-AAGIGILTV poorly. Structural analysis revealed the potential for new contacts between the MEL5 T-cell receptor and the optimized peptide. Furthermore, the optimized peptide was able to prime CD8(+) T-cell populations in peripheral blood mononuclear cell isolates from multiple HLA A*0201(+) individuals that were capable of efficient HLA A*0201(+) melanoma cell destruction. This proof-of-concept study demonstrates that it is possible to design altered peptide antigens for the selection of superior T-cell clonotypes with enhanced antigen recognition properties.
- Institute of Infection and Immunity, Cardiff University School of Medicine, Henry Wellcome Building, Heath Park, Cardiff CF14 4XN, Wales, United Kingdom.
Organizational Affiliation: