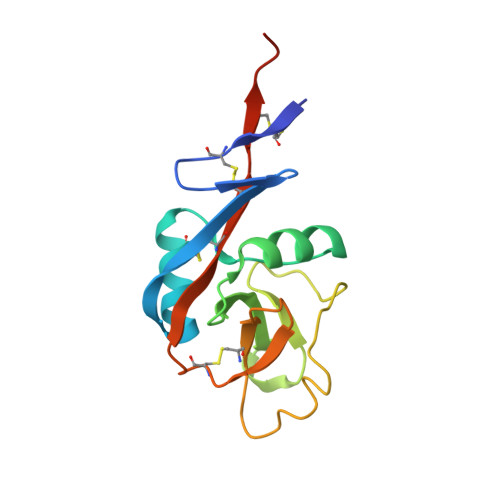
Ca2+-dependent Structural Changes in the B-cell Receptor CD23 Increase Its Affinity for Human Immunoglobulin E.
Yuan, D., Keeble, A.H., Hibbert, R.G., Fabiane, S., Gould, H.J., McDonnell, J.M., Beavil, A.J., Sutton, B.J., Dhaliwal, B.(2013) J Biological Chem 288: 21667-21677
- PubMed: 23775083 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.480657
- Primary Citation Related Structures:
4G96, 4G9A, 4GI0, 4GJ0, 4GJX, 4GK1, 4GKO - PubMed Abstract:
Immunoglobulin E (IgE) antibodies play a fundamental role in allergic disease and are a target for therapeutic intervention. IgE functions principally through two receptors, FcεRI and CD23 (FcεRII). Minute amounts of allergen trigger mast cell or basophil degranulation by cross-linking IgE-bound FcεRI, leading to an inflammatory response. The interaction between IgE and CD23 on B-cells regulates IgE synthesis. CD23 is unique among Ig receptors in that it belongs to the C-type (calcium-dependent) lectin-like superfamily. Although the interaction of CD23 with IgE is carbohydrate-independent, calcium has been reported to increase the affinity for IgE, but the structural basis for this activity has previously been unknown. We have determined the crystal structures of the human lectin-like head domain of CD23 in its Ca(2+)-free and Ca(2+)-bound forms, as well as the crystal structure of the Ca(2+)-bound head domain of CD23 in complex with a subfragment of IgE-Fc consisting of the dimer of Cε3 and Cε4 domains (Fcε3-4). Together with site-directed mutagenesis, the crystal structures of four Ca(2+) ligand mutants, isothermal titration calorimetry, surface plasmon resonance, and stopped-flow analysis, we demonstrate that Ca(2+) binds at the principal and evolutionarily conserved binding site in CD23. Ca(2+) binding drives Pro-250, at the base of an IgE-binding loop (loop 4), from the trans to the cis configuration with a concomitant conformational change and ordering of residues in the loop. These Ca(2+)-induced structural changes in CD23 lead to additional interactions with IgE, a more entropically favorable interaction, and a 30-fold increase in affinity of a single head domain of CD23 for IgE. Taken together, these results suggest that binding of Ca(2+) brings an extra degree of modulation to CD23 function.
- King's College London and the Medical Research Council and Asthma UK Centre in Allergic Mechanisms of Asthma, Randall Division of Cell and Molecular Biophysics, Guy's Campus, London, SE1 1UL, United Kingdom.
Organizational Affiliation: