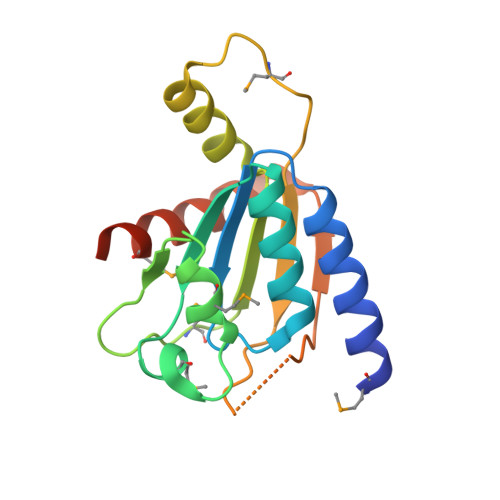
Structure and function of Zucchini endoribonuclease in piRNA biogenesis
Nishimasu, H., Ishizu, H., Saito, K., Fukuhara, S., Kamatani, M.K., Bonnefond, L., Matsumoto, N., Nishizawa, T., Nakanaga, K., Aoki, J., Ishitani, R., Siomi, H., Siomi, M.C., Nureki, O.(2012) Nature 491: 284-287
- PubMed: 23064230 Search on PubMed
- DOI: https://doi.org/10.1038/nature11509
- Primary Citation Related Structures:
4GEL, 4GEM, 4GEN - PubMed Abstract:
PIWI-interacting RNAs (piRNAs) silence transposons to maintain genome integrity in animal germ lines. piRNAs are classified as primary and secondary piRNAs, depending on their biogenesis machinery. Primary piRNAs are processed from long non-coding RNA precursors transcribed from piRNA clusters in the genome through the primary processing pathway. Although the existence of a ribonuclease participating in this pathway has been predicted, its molecular identity remained unknown. Here we show that Zucchini (Zuc), a mitochondrial phospholipase D (PLD) superfamily member, is an endoribonuclease essential for primary piRNA biogenesis. We solved the crystal structure of Drosophila melanogaster Zuc (DmZuc) at 1.75 Å resolution. The structure revealed that DmZuc has a positively charged, narrow catalytic groove at the dimer interface, which could accommodate a single-stranded, but not a double-stranded, RNA. DmZuc and the mouse homologue MmZuc (also known as Pld6 and MitoPLD) showed endoribonuclease activity for single-stranded RNAs in vitro. The RNA cleavage products bear a 5'-monophosphate group, a hallmark of mature piRNAs. Mutational analyses revealed that the conserved active-site residues of DmZuc are critical for the ribonuclease activity in vitro, and for piRNA maturation and transposon silencing in vivo. We propose a model for piRNA biogenesis in animal germ lines, in which the Zuc endoribonuclease has a key role in primary piRNA maturation.
- Department of Biophysics and Biochemistry, Graduate School of Science, The University of Tokyo, Tokyo 113-0032, Japan.
Organizational Affiliation: