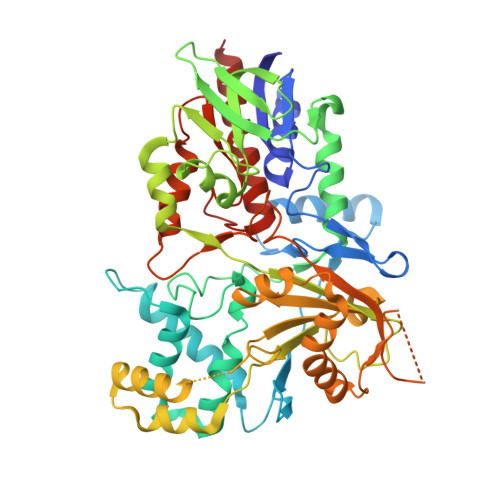
Mechanistic and Structural Analyses of the Roles of Active Site Residues in Yeast Polyamine Oxidase Fms1: Characterization of the N195A and D94N Enzymes.
Adachi, M.S., Taylor, A.B., Hart, P.J., Fitzpatrick, P.F.(2012) Biochemistry 51: 8690-8697
- PubMed: 23034052 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi3011434
- Primary Citation Related Structures:
4GDP - PubMed Abstract:
Flavoprotein Fms1 from Saccharomyces cerevisiae catalyzes the oxidation of spermine in the biosynthetic pathway for pantothenic acid. The same reaction is catalyzed by the mammalian polyamine and spermine oxidases. The active site of Fms1 contains three amino acid residues positioned to interact with the polyamine substrate, His67, Asn195, and Asp94. These three residues form a hydrogen-bonding triad with Asn195 being the central residue. Previous studies of the effects of mutating His67 are consistent with that residue being important both for interacting with the substrate and for maintaining the hydrogen bonds in the triad [Adachi, M. S., Taylor, A. B., Hart, P. J., and Fitzpatrick, P. F. (2012) Biochemistry 51, 4888-4897]. The N195A and D94N enzymes have now been characterized to evaluate their roles in catalysis. Both mutations primarily affect the reductive half-reaction. With N(1)-acetylspermine as the substrate, the rate constant for flavin reduction decreases ~450-fold for both mutations; the effects with spermine as the substrate are smaller, 20-40-fold. The k(cat)/K(amine)- and k(cat)-pH profiles with N(1)-acetylspermine are only slightly changed from the profiles for the wild-type enzyme, consistent with the pK(a) values arising from the amine substrate or product and not from active site residues. The structure of the N195A enzyme was determined at a resolution of 2.0 Å. The structure shows a molecule of tetraethylene glycol in the active site and establishes that the mutation has no effect on the protein structure. Overall, the results are consistent with the role of Asn195 and Asp94 being to properly position the polyamine substrate for oxidation.
- Department of Biochemistry, University of Texas Health Science Center at San Antonio, San Antonio, TX 78229, USA.
Organizational Affiliation: