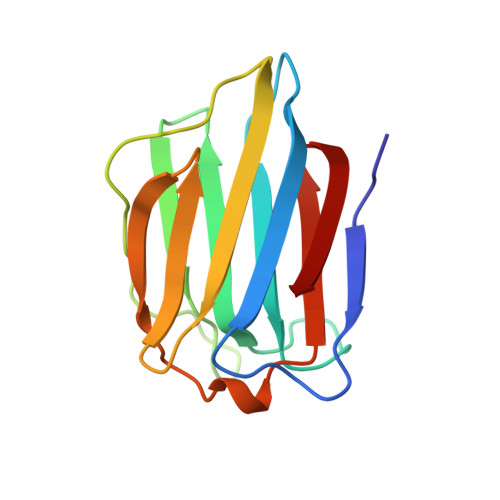
Structural basis for the recognition of carbohydrates by human galectin-7.
Leonidas, D.D., Vatzaki, E.H., Vorum, H., Celis, J.E., Madsen, P., Acharya, K.R.(1998) Biochemistry 37: 13930-13940
- PubMed: 9760227 Search on PubMed
- DOI: https://doi.org/10.1021/bi981056x
- Primary Citation Related Structures:
1BKZ, 2GAL, 3GAL, 4GAL, 5GAL - PubMed Abstract:
Knowledge about carbohydrate recognition domains of galectins, formerly known as S-type animal lectins, is important in understanding their role(s) in cell-cell interactions. Here we report the crystal structure of human galectin-7 (hGal-7), in free form and in the presence of galactose, galactosamine, lactose, and N-acetyl-lactosamine at high resolution. This is the first structure of a galectin determined in both free and carbohydrate-bound forms. The structure shows a fold similar to that of the prototype galectins -1 and -2, but has greater similarity to a related galectin molecule, Gal-10. Even though the carbohydrate-binding residues are conserved, there are significant changes in this pocket due to shortening of a loop structure. The monomeric hGal-7 molecule exists as a dimer in the crystals, but adopts a packing arrangement considerably different from that of Gal-1 and Gal-2, which has implications for carbohydrate recognition.
- Department of Biology and Biochemistry, University of Bath, UK.
Organizational Affiliation: