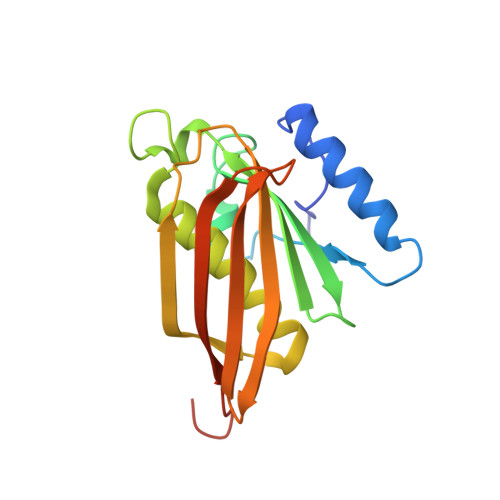
Correlation of structure and function in the human hotdog-fold enzyme hTHEM4.
Zhao, H., Lim, K., Choudry, A., Latham, J.A., Pathak, M.C., Dominguez, D., Luo, L., Herzberg, O., Dunaway-Mariano, D.(2012) Biochemistry 51: 6490-6492
- PubMed: 22871024 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi300968n
- Primary Citation Related Structures:
4GAH - PubMed Abstract:
Human THEM4 (hTHEM4) is comprised of a catalytically active hotdog-fold acyl-CoA thioesterase domain and an N-terminal domain of unknown fold and function. hTHEM4 has been linked to Akt1 regulation and cell apoptosis. Herein, we report the X-ray structure of hHTEM4 bound with undecan-2-one-CoA. Structure guided mutagenesis was carried out to confirm the catalytic residues. The N-terminal domain is shown to be partially comprised of irregular and flexible secondary structure, reminiscent of a protein-binding domain. We demonstrate direct hTHEM4-Akt1 binding by immunoprecipitation and by inhibition of Akt1 kinase activity, thus providing independent evidence that hTHEM4 is an Akt1 negative regulator.
- Department of Chemistry and Chemical Biology, University of New Mexico, Albuquerque, NM 87131, USA.
Organizational Affiliation: