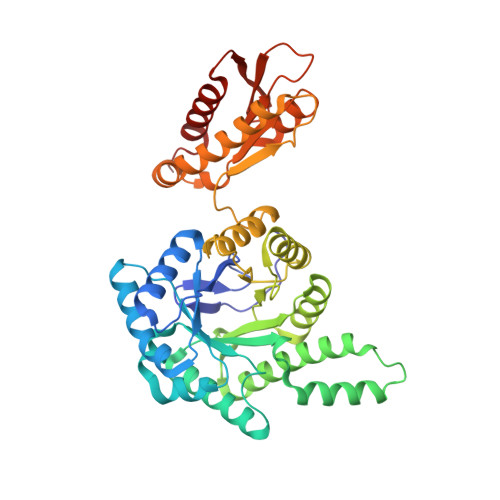
Structure of the GcpE (IspG)-MEcPP complex from Thermus thermophilus.
Rekittke, I., Jomaa, H., Ermler, U.(2012) FEBS Lett 586: 3452-3457
- PubMed: 22967895 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2012.07.070
- Primary Citation Related Structures:
4G9P - PubMed Abstract:
Isoprenoid precursor biosynthesis occurs through the mevalonate or the methylerythritol phosphate (MEP) pathway, used i.e., by humans and by many human pathogens, respectively. In the MEP pathway, 2-C-methyl-D-erythritol-2,4-cyclo-diphosphate (MEcPP) is converted to (E)-1-hydroxy-2-methyl-but-2-enyl-4-diphosphate (HMBPP) by the iron-sulfur cluster enzyme HMBPP synthase (GcpE). The presented X-ray structure of the GcpE-MEcPP complex from Thermus thermophilus at 1.55Å resolution provides valuable information about the catalytic mechanism and for rational inhibitor design. MEcPP binding inside the TIM-barrel funnel induces a 60° rotation of the [4Fe-4S] cluster containing domain onto the TIM-barrel entrance. The apical iron of the [4Fe-4S] cluster ligates with the C3 oxygen atom of MEcPP.
- Institut für Klinische Immunologie und Transfusionsmedizin, Justus-Liebig-Universität Giessen, Giessen, Germany.
Organizational Affiliation: