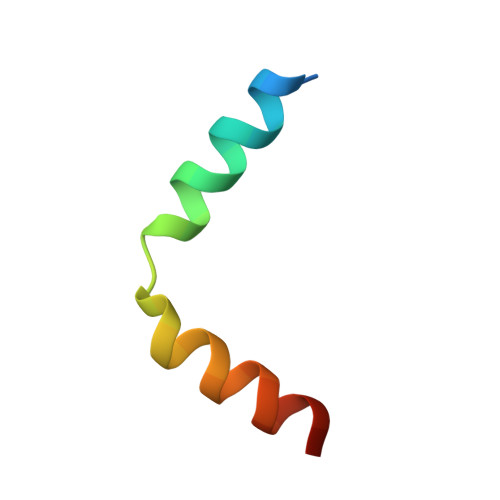
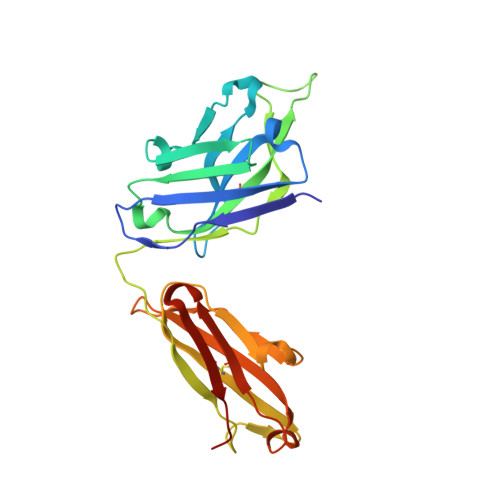
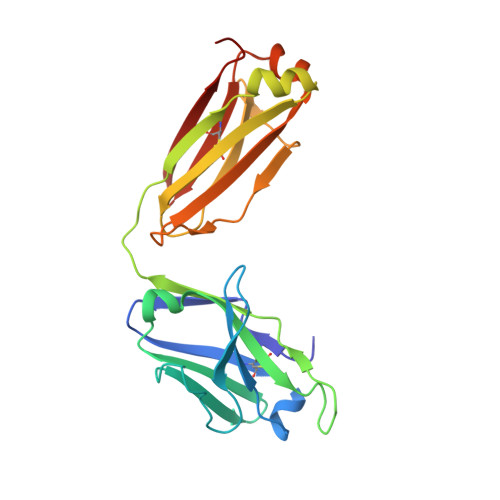
Broad and potent neutralization of HIV-1 by a gp41-specific human antibody.
Huang, J., Ofek, G., Laub, L., Louder, M.K., Doria-Rose, N.A., Longo, N.S., Imamichi, H., Bailer, R.T., Chakrabarti, B., Sharma, S.K., Alam, S.M., Wang, T., Yang, Y., Zhang, B., Migueles, S.A., Wyatt, R., Haynes, B.F., Kwong, P.D., Mascola, J.R., Connors, M.(2012) Nature 491: 406-412
- PubMed: 23151583 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature11544
- Primary Citation Related Structures:
4G6F - PubMed Abstract:
Characterization of human monoclonal antibodies is providing considerable insight into mechanisms of broad HIV-1 neutralization. Here we report an HIV-1 gp41 membrane-proximal external region (MPER)-specific antibody, named 10E8, which neutralizes ∼98% of tested viruses. An analysis of sera from 78 healthy HIV-1-infected donors demonstrated that 27% contained MPER-specific antibodies and 8% contained 10E8-like specificities. In contrast to other neutralizing MPER antibodies, 10E8 did not bind phospholipids, was not autoreactive, and bound cell-surface envelope. The structure of 10E8 in complex with the complete MPER revealed a site of vulnerability comprising a narrow stretch of highly conserved gp41-hydrophobic residues and a critical arginine or lysine just before the transmembrane region. Analysis of resistant HIV-1 variants confirmed the importance of these residues for neutralization. The highly conserved MPER is a target of potent, non-self-reactive neutralizing antibodies, suggesting that HIV-1 vaccines should aim to induce antibodies to this region of HIV-1 envelope glycoprotein.
- HIV-Specific Immunity Section, Laboratory of Immunoregulation, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, Maryland 20892, USA.
Organizational Affiliation: