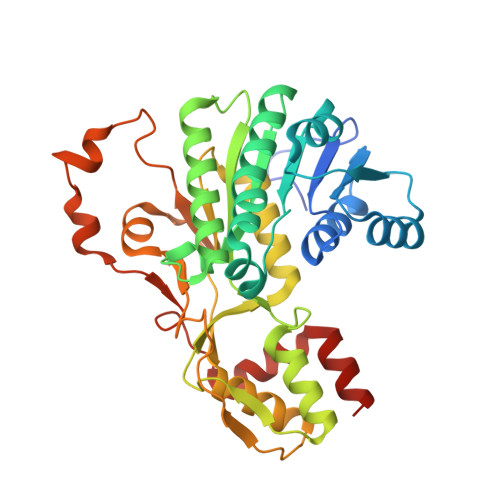
Dynamic elements govern the catalytic activity of CapE, a capsular polysaccharide-synthesizing enzyme from Staphylococcus aureus.
Miyafusa, T., Caaveiro, J.M., Tanaka, Y., Tsumoto, K.(2013) FEBS Lett 587: 3824-3830
- PubMed: 24157361 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2013.10.009
- Primary Citation Related Structures:
4G5H - PubMed Abstract:
CapE is an essential enzyme for the synthesis of capsular polysaccharide (CP) of pathogenic strains of Staphylococcus aureus. Herein we demonstrate that CapE is a 5-inverting 4,6-dehydratase enzyme. However, in the absence of downstream enzymes, CapE catalyzes an additional reaction (5-back-epimerization) affording a by-product under thermodynamic control. Single-crystal X-ray crystallography was employed to identify the structure of the by-product. The structural analysis reveals a network of coordinated motions away from the active site governing the enzymatic activity of CapE. A second dynamic element (the latch) regulates the enzymatic chemoselectivity. The validity of these mechanisms was evaluated by site-directed mutagenesis.
- Medical Proteomics Laboratory, Institute of Medical Science, The University of Tokyo, Minato-ku, Tokyo 108-8639, Japan; Department of Medical Genome Sciences, School of Frontier Sciences, The University of Tokyo, Minato-ku, Tokyo 108-8639, Japan.
Organizational Affiliation: