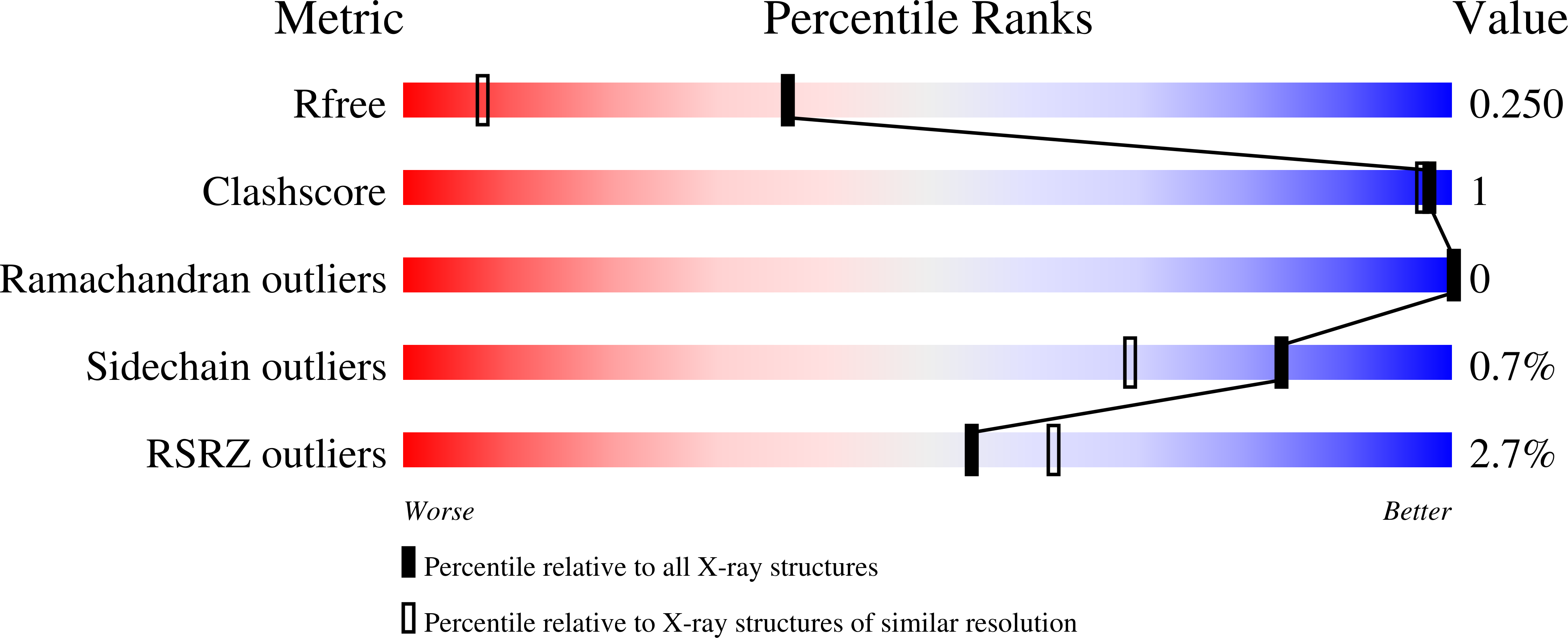
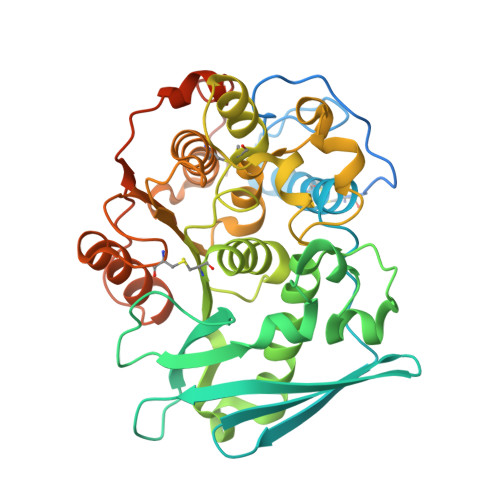
The structure of a novel glucuronoyl esterase from Myceliophthora thermophila gives new insights into its role as a potential biocatalyst.
Charavgi, M.D., Dimarogona, M., Topakas, E., Christakopoulos, P., Chrysina, E.D.(2013) Acta Crystallogr D Biol Crystallogr 69: 63-73
- PubMed: 23275164
- DOI: https://doi.org/10.1107/S0907444912042400
- Primary Citation of Related Structures:
4G4G, 4G4I, 4G4J - PubMed Abstract:
The increasing demand for the development of efficient biocatalysts is a consequence of their broad industrial applications. Typical difficulties that are encountered during their exploitation in a variety of processes are interconnected with factors such as temperature, pH, product inhibitors etc. To eliminate these, research has been directed towards the identification of new enzymes that would comply with the required standards. To this end, the recently discovered glucuronoyl esterases (GEs) are an enigmatic family within the carbohydrate esterase (CE) family. Structures of the thermophilic StGE2 esterase from Myceliophthora thermophila (synonym Sporotrichum thermophile), a member of the CE15 family, and its S213A mutant were determined at 1.55 and 1.9 Å resolution, respectively. The first crystal structure of the S213A mutant in complex with a substrate analogue, methyl 4-O-methyl-β-D-glucopyranuronate, was determined at 2.35 Å resolution. All of the three-dimensional protein structures have an α/β-hydrolase fold with a three-layer αβα-sandwich architecture and a Rossmann topology and comprise one molecule per asymmetric unit. These are the first crystal structures of a thermophilic GE both in an unliganded form and bound to a substrate analogue, thus unravelling the organization of the catalytic triad residues and their neighbours lining the active site. The knowledge derived offers novel insights into the key structural elements that drive the hydrolysis of glucuronic acid esters.
Organizational Affiliation:
Institute of Biology, Medicinal Chemistry and Biotechnology, National Hellenic Research Foundation, Athens, Greece.