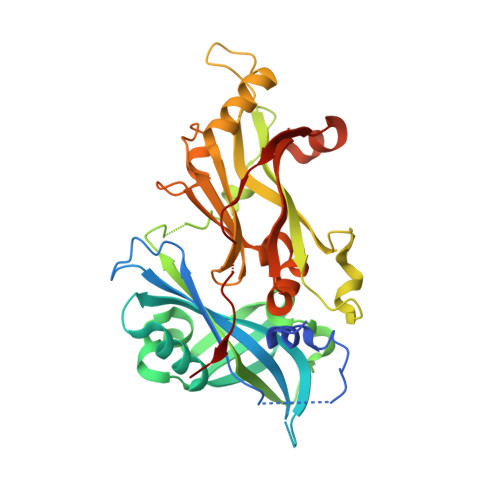
Structure and assembly of a paramyxovirus matrix protein.
Battisti, A.J., Meng, G., Winkler, D.C., McGinnes, L.W., Plevka, P., Steven, A.C., Morrison, T.G., Rossmann, M.G.(2012) Proc Natl Acad Sci U S A 109: 13996-14000
- PubMed: 22891297 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1210275109
- Primary Citation Related Structures:
4G1G, 4G1L, 4G1O - PubMed Abstract:
Many pleomorphic, lipid-enveloped viruses encode matrix proteins that direct their assembly and budding, but the mechanism of this process is unclear. We have combined X-ray crystallography and cryoelectron tomography to show that the matrix protein of Newcastle disease virus, a paramyxovirus and relative of measles virus, forms dimers that assemble into pseudotetrameric arrays that generate the membrane curvature necessary for virus budding. We show that the glycoproteins are anchored in the gaps between the matrix proteins and that the helical nucleocapsids are associated in register with the matrix arrays. About 90% of virions lack matrix arrays, suggesting that, in agreement with previous biological observations, the matrix protein needs to dissociate from the viral membrane during maturation, as is required for fusion and release of the nucleocapsid into the host's cytoplasm. Structure and sequence conservation imply that other paramyxovirus matrix proteins function similarly.
- Department of Biological Sciences, Purdue University, West Lafayette, IN 47907-2032, USA.
Organizational Affiliation: