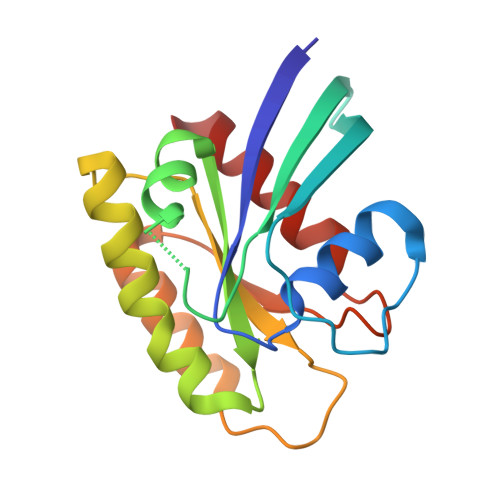
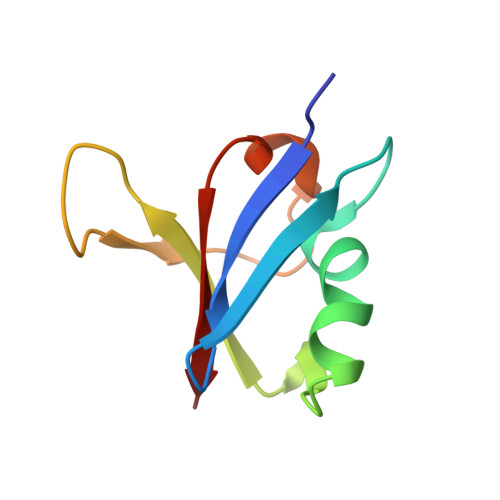
Allosteric Effects of the Oncogenic RasQ61L Mutant on Raf-RBD.
Fetics, S.K., Guterres, H., Kearney, B.M., Buhrman, G., Ma, B., Nussinov, R., Mattos, C.(2015) Structure 23: 505-516
- PubMed: 25684575 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2014.12.017
- Primary Citation Related Structures:
4G0N, 4G3X - PubMed Abstract:
The Ras/Raf/MEK/ERK signal transduction pathway is a major regulator of cell proliferation activated by Ras-guanosine triphosphate (GTP). The oncogenic mutant RasQ61L is not able to hydrolyze GTP in the presence of Raf and thus is a constitutive activator of this mitogenic pathway. The Ras/Raf interaction is essential for the activation of the Raf kinase domain through a currently unknown mechanism. We present the crystal structures of the Ras-GppNHp/Raf-RBD and RasQ61L-GppNHp/Raf-RBD complexes, which, in combination with MD simulations, reveal differences in allosteric interactions leading from the Ras/Raf interface to the Ras calcium-binding site and to the remote Raf-RBD loop L4. In the presence of Raf, the RasQ61L mutant has a rigid switch II relative to the wild-type and increased flexibility at the interface with switch I, which propagates across Raf-RBD. We show that in addition to local perturbations on Ras, RasQ61L has substantial long-range effects on the Ras allosteric lobe and on Raf-RBD.
- Department of Chemistry and Chemical Biology, Northeastern University, Boston, MA 02115, USA; Department of Molecular and Structural Biochemistry, North Carolina State University, Raleigh, NC 27695, USA.
Organizational Affiliation: