Discovery and Mechanism Study of SIRT1 Activators that Promote the Deacetylation of Fluorophore-Labeled Substrate
Wu, J., Zhang, D., Chen, L., Li, J., Wang, J., Ning, C., Yu, N., Zhao, F., Chen, D., Chen, X., Chen, K., Jiang, H., Liu, H., Liu, D.(2013) J Med Chem 56: 761-780
- PubMed: 23316803 Search on PubMed
- DOI: https://doi.org/10.1021/jm301032j
- Primary Citation Related Structures:
4FZ3 - PubMed Abstract:
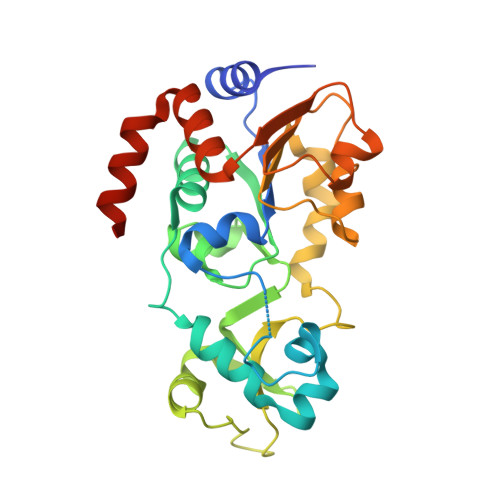
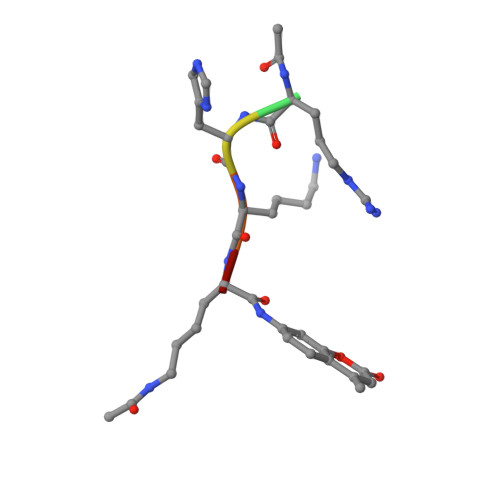
SIRT1 is an NAD(+)-dependent deacetylase, whose activators have potential therapeutic applications in age-related diseases. Here we report a new class of SIRT1 activators. The activation is dependent on the fluorophore labeled to the substrate. To elucidate the activation mechanism, we solved the crystal structure of SIRT3/ac-RHKK(ac)-AMC complex. The structure revealed that the fluorophore blocked the H-bond formation and created a cavity between the substrate and the Rossmann fold. We built the SIRT1/ac-RHKK(ac)-AMC complex model based on the crystal structure. K(m) and K(d) determinations demonstrated that the fluorophore decreased the peptide binding affinity. The binding modes of SIRT1 activators indicated that a portion of the activators interacts with the fluorophore through π-stacking, while the other portion inserts into the cavity or interacts with the Rossmann fold, thus increasing the substrate affinity. Our study provides new insights into the mechanism of SIRT1 activation and may aid the design of novel SIRT1 activators.
- Department of Pharmacology III, Shanghai Institute of Materia Medica , Chinese Academy of Sciences, 555 Zu-Chong-Zhi Road, Shanghai 201203, China.
Organizational Affiliation: