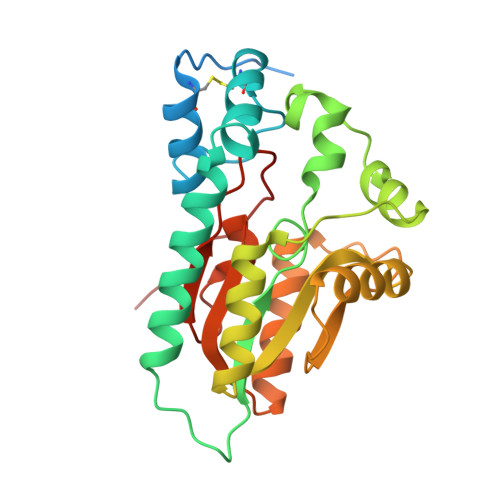
The crystal structure of Arabidopsis VSP1 reveals the plant class C-like phosphatase structure of the DDDD superfamily of phosphohydrolases
Chen, Y., Wei, J., Wang, M., Shi, Z., Gong, W., Zhang, M.(2012) PLoS One 7: e49421-e49421
- PubMed: 23166664 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0049421
- Primary Citation Related Structures:
4FYP - PubMed Abstract:
Arabidopsis thaliana vegetative storage proteins, VSP1 and VSP2, are acid phosphatases and belong to the haloacid dehalogenase (HAD) superfamily. In addition to their potential nutrient storage function, they were thought to be involved in plant defense and flower development. To gain insights into the architecture of the protein and obtain clues about its function, we have tested their substrate specificity and solved the structure of VSP1. The acid phosphatase activities of these two enzymes require divalent metal such as magnesium ion. Conversely, the activity of these two enzymes is inhibited by vanadate and molybdate, but is resistant to inorganic phosphate. Both VSP1 and VSP2 did not exhibit remarkable activities to any physiological substrates tested. In the current study, we presented the crystal structure of recombinant VSP1 at 1.8 Å resolution via the selenomethionine single-wavelength anomalous diffraction (SAD). Specifically, an α-helical cap domain on the top of the α/β core domain is found to be involved in dimerization. In addition, despite of the low sequence similarity between VSP1 and other HAD enzymes, the core domain of VSP1 containing conserved active site and catalytic machinery displays a classic haloacid dehalogenase fold. Furthermore, we found that VSP1 is distinguished from bacterial class C acid phosphatase P4 by several structural features. To our knowledge, this is the first study to reveal the crystal structure of plant vegetative storage proteins.
- School of Life Sciences, Anhui University, Hefei, Anhui, China.
Organizational Affiliation: