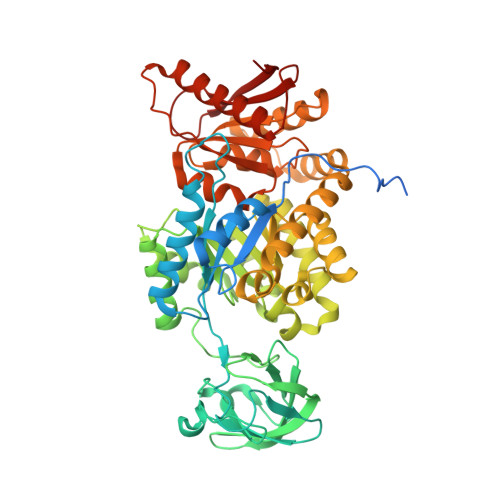
M2 pyruvate kinase provides a mechanism for nutrient sensing and regulation of cell proliferation.
Morgan, H.P., O'Reilly, F.J., Wear, M.A., O'Neill, J.R., Fothergill-Gilmore, L.A., Hupp, T., Walkinshaw, M.D.(2013) Proc Natl Acad Sci U S A 110: 5881-5886
- PubMed: 23530218 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1217157110
- Primary Citation Related Structures:
4FXF, 4FXJ - PubMed Abstract:
We show that the M2 isoform of pyruvate kinase (M2PYK) exists in equilibrium between monomers and tetramers regulated by allosteric binding of naturally occurring small-molecule metabolites. Phenylalanine stabilizes an inactive T-state tetrameric conformer and inhibits M2PYK with an IC50 value of 0.24 mM, whereas thyroid hormone (triiodo-L-thyronine, T3) stabilizes an inactive monomeric form of M2PYK with an IC50 of 78 nM. The allosteric activator fructose-1,6-bisphosphate [F16BP, AC50 (concentration that gives 50% activation) of 7 μM] shifts the equilibrium to the tetrameric active R-state, which has a similar activity to that of the constitutively fully active isoform M1PYK. Proliferation assays using HCT-116 cells showed that addition of inhibitors phenylalanine and T3 both increased cell proliferation, whereas addition of the activator F16BP reduced proliferation. F16BP abrogates the inhibitory effect of both phenylalanine and T3, highlighting a dominant role of M2PYK allosteric activation in the regulation of cancer proliferation. X-ray structures show constitutively fully active M1PYK and F16BP-bound M2PYK in an R-state conformation with a lysine at the dimer-interface acting as a peg in a hole, locking the active tetramer conformation. Binding of phenylalanine in an allosteric pocket induces a 13° rotation of the protomers, destroying the peg-in-hole R-state interface. This distinct T-state tetramer is stabilized by flipped out Trp/Arg side chains that stack across the dimer interface. X-ray structures and biophysical binding data of M2PYK complexes explain how, at a molecular level, fluctuations in concentrations of amino acids, thyroid hormone, and glucose metabolites switch M2PYK on and off to provide the cell with a nutrient sensing and growth signaling mechanism.
- Centre for Translational and Chemical Biology, University of Edinburgh, Edinburgh EH9 3JR, United Kingdom.
Organizational Affiliation: