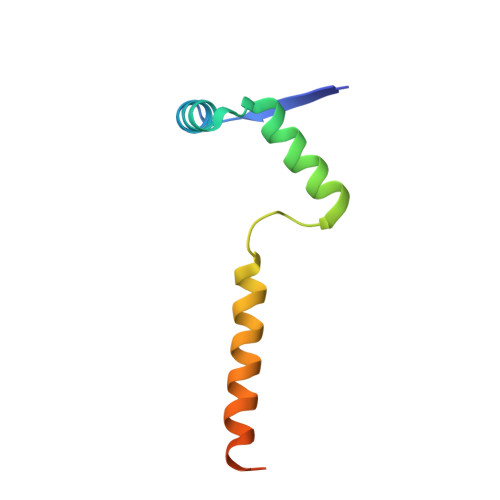
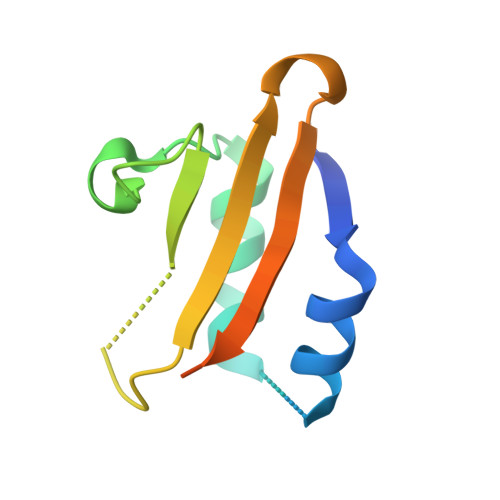
The crystal structure of the intact E. coli RelBE toxin-antitoxin complex provides the structural basis for conditional cooperativity.
Boggild, A., Sofos, N., Andersen, K.R., Feddersen, A., Easter, A.D., Passmore, L.A., Brodersen, D.E.(2012) Structure 20: 1641-1648
- PubMed: 22981948 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2012.08.017
- Primary Citation Related Structures:
4FXE, 4FXH, 4FXI - PubMed Abstract:
The bacterial relBE locus encodes a toxin-antitoxin complex in which the toxin, RelE, is capable of cleaving mRNA in the ribosomal A site cotranslationally. The antitoxin, RelB, both binds and inhibits RelE, and regulates transcription through operator binding and conditional cooperativity controlled by RelE. Here, we present the crystal structure of the intact Escherichia coli RelB2E2 complex at 2.8 Å resolution, comprising both the RelB-inhibited RelE and the RelB dimerization domain that binds DNA. RelE and RelB associate into a V-shaped heterotetrameric complex with the ribbon-helix-helix (RHH) dimerization domain at the apex. Our structure supports a model in which relO is optimally bound by two adjacent RelB2E heterotrimeric units, and is not compatible with concomitant binding of two RelB2E2 heterotetramers. The results thus provide a firm basis for understanding the model of conditional cooperativity at the molecular level.
- Centre for mRNP Biogenesis and Metabolism, Department of Molecular Biology and Genetics, Aarhus University, DK-8000 Aarhus C, Denmark.
Organizational Affiliation: