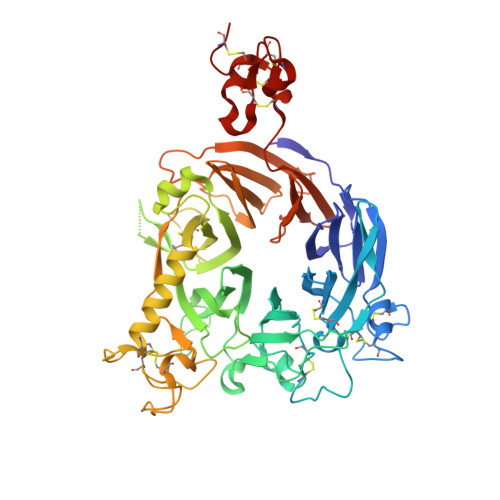
Crystal Structure of the Sema-PSI Extracellular Domain of Human RON Receptor Tyrosine Kinase.
Chao, K.L., Tsai, I.W., Chen, C., Herzberg, O.(2012) PLoS One 7: e41912-e41912
- PubMed: 22848655 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0041912
- Primary Citation Related Structures:
4FWW - PubMed Abstract:
Human RON (Recepteur d'Origine Nantais) receptor tyrosine kinase is a cell surface receptor for Macrophage Stimulating Protein (MSP). RON mediates signal transduction pathways that regulate cell adhesion, invasion, motility and apoptosis processes. Elevated levels of RON and its alternatively spliced variants are implicated in the progression and metastasis of tumor cells. The binding of MSP α/β heterodimer to the extracellular region of RON receptor induces receptor dimerization and activation by autophosphorylation of the intracellular kinase domains. The ectodomain of RON, containing the ligand recognition and dimerization domains, is composed of a semaphorin (Sema), Plexins-Semaphorins-Integrins domain (PSI), and four Immunoglobulins-Plexins-Transcription factor (IPT) domains. High affinity association between MSP and RON is mediated by the interaction between MSP β-chain and RON Sema, although RON activation requires intact RON and MSP proteins. Here, we report the structure of RON Sema-PSI domains at 1.85 Å resolution. RON Sema domain adopts a seven-bladed β-propeller fold, followed by disulfide bond rich, cysteine-knot PSI motif. Comparison with the homologous Met receptor tyrosine kinase reveals that RON Sema-PSI contains distinguishing secondary structural features. These define the receptors' exclusive selectivity towards their respective ligands, RON for MSP and Met for HGF. The RON Sema-PSI crystal packing generates a homodimer with interface formed by the Sema domain. Mapping of the dimer interface using the RON homology to Met, MSP homology to Hepatocyte Growth Factor (HGF), and the structure of the Met/HGF complex shows the dimer interface overlapping with the putative MSPβ binding site. The crystallographically determined RON Sema-PSI homodimer may represent the dimer assembly that occurs during ligand-independent receptor activation and/or the inhibition of the constitutive activity of RONΔ160 splice variant by the soluble RON splice variant, RONΔ85.
- Institute for Bioscience and Biotechnology Research, University of Maryland, Rockville, Maryland, United States of America.
Organizational Affiliation: