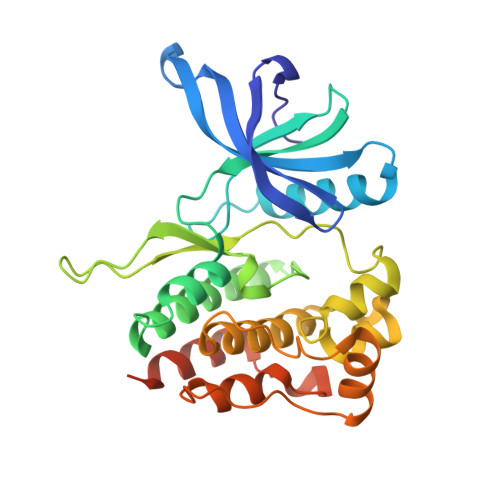
Crystal structures of the JAK2 pseudokinase domain and the pathogenic mutant V617F.
Bandaranayake, R.M., Ungureanu, D., Shan, Y., Shaw, D.E., Silvennoinen, O., Hubbard, S.R.(2012) Nat Struct Mol Biol 19: 754-759
- PubMed: 22820988 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2348
- Primary Citation Related Structures:
4FVP, 4FVQ, 4FVR - PubMed Abstract:
The protein tyrosine kinase JAK2 mediates signaling through numerous cytokine receptors. JAK2 possesses a pseudokinase domain (JH2) and a tyrosine kinase domain (JH1). Through unknown mechanisms, JH2 regulates the catalytic activity of JH1, and hyperactivating mutations in the JH2 region of human JAK2 cause myeloproliferative neoplasms (MPNs). We showed previously that JAK2 JH2 is, in fact, catalytically active. Here we present crystal structures of human JAK2 JH2, including both wild type and the most prevalent MPN mutant, V617F. The structures reveal that JH2 adopts the fold of a prototypical protein kinase but binds Mg-ATP noncanonically. The structural and biochemical data indicate that the V617F mutation rigidifies α-helix C in the N lobe of JH2, facilitating trans-phosphorylation of JH1. The crystal structures of JH2 afford new opportunities for the design of novel JAK2 therapeutics targeting MPNs.
- Structural Biology Program, Kimmel Center for Biology and Medicine at the Skirball Institute, New York University School of Medicine, New York, New York, USA.
Organizational Affiliation: