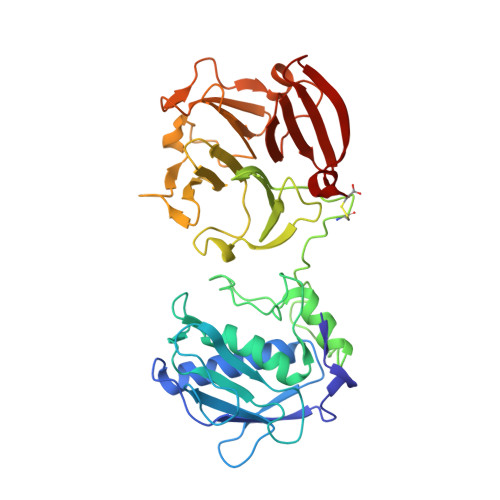
Crystal structure of full-length human collagenase 3 (MMP-13) with peptides in the active site defines exosites in the catalytic domain.
Stura, E.A., Visse, R., Cuniasse, P., Dive, V., Nagase, H.(2013) FASEB J 27: 4395-4405
- PubMed: 23913860 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1096/fj.13-233601
- Primary Citation Related Structures:
4FU4, 4FVL, 4G0D - PubMed Abstract:
Matrix metalloproteinase (MMP)-13 is one of the mammalian collagenases that play key roles in tissue remodelling and repair and in progression of diseases such as cancer, arthritis, atherosclerosis, and aneurysm. For collagenase to cleave triple helical collagens, the triple helical structure has to be locally unwound before hydrolysis, but this process is not well understood. We report crystal structures of catalytically inactive full-length human MMP-13(E223A) in complex with peptides of 14-26 aa derived from the cleaved prodomain during activation. Peptides are bound to the active site of the enzyme by forming an extended β-strand with Glu(40) or Tyr(46) inserted into the S1' specificity pocket. The structure of the N-terminal part of the peptides is variable and interacts with different parts of the catalytic domain. Those areas are designated substrate-dependent exosites, in that they accommodate different peptide structures, whereas the precise positioning of the substrate backbone is maintained in the active site. These modes of peptide-MMP-13 interactions have led us to propose how triple helical collagen strands fit into the active site cleft of the collagenase.
- 2H.N., Kennedy Institute of Rheumatology, Nuffield Department of Orthopaedics, Rheumatology, and Musculoskeletal Sciences, University of Oxford, Roosevelt Drive, Oxford, OX3 7FY, UK. hideaki.nagase@kennedy.ox.ac.uk.
Organizational Affiliation: