Structure of a high affinity TCR reveals native binding to MHC with enhanced peptide specificity
Borbulevych, O.Y., Baker, B.M.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| HLA class I histocompatibility antigen, A-2 alpha chain | 275 | Homo sapiens | Mutation(s): 0 Gene Names: HLA-A, HLAA | 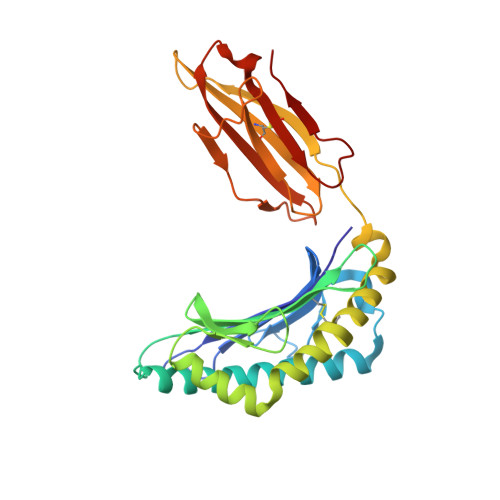 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P04439 GTEx: ENSG00000206503 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P04439 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Beta-2-microglobulin | 100 | Homo sapiens | Mutation(s): 0 Gene Names: B2M, CDABP0092, HDCMA22P | 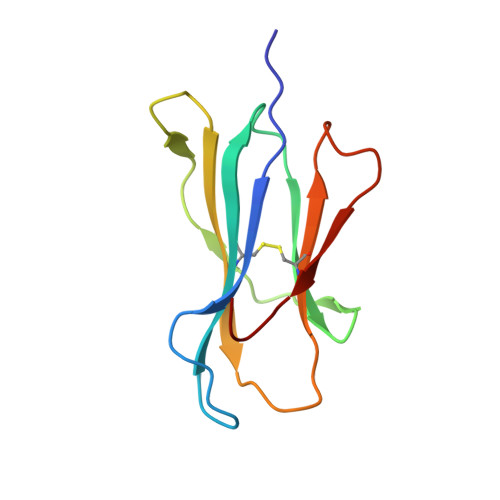 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P61769 GTEx: ENSG00000166710 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P61769 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Protein Tax-1 | 9 | Human T-cell lymphotropic virus type 1 (african isolate) | Mutation(s): 0 | 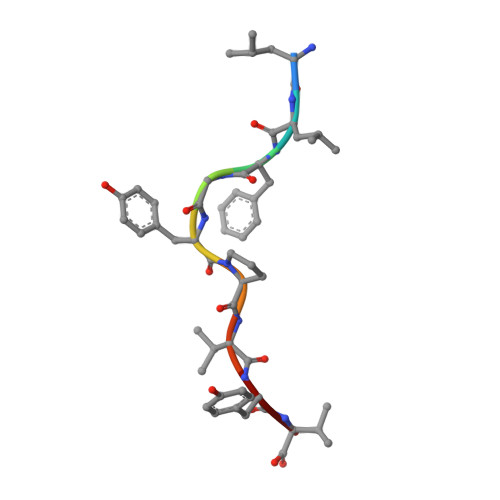 | |
UniProt | |||||
Entity Groups | |||||
| UniProt Group | P0C213 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| A6 alpha chain | 200 | Homo sapiens | Mutation(s): 0 | 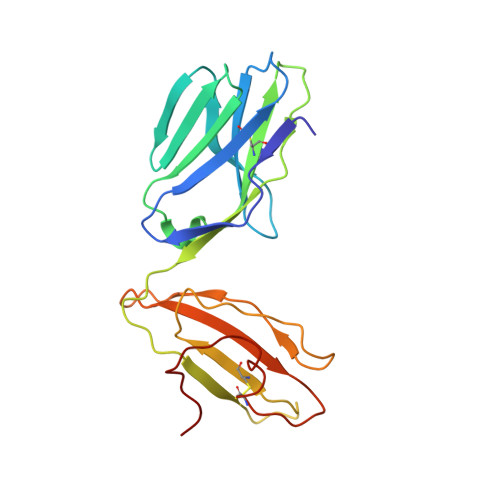 | |
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| A6 beta chain | 245 | Homo sapiens | Mutation(s): 0 |  | |
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| GOL Download:Ideal Coordinates CCD File | F [auth A] G [auth A] H [auth A] I [auth A] J [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 224.23 | α = 90 |
| b = 48.954 | β = 89.62 |
| c = 94.398 | γ = 90 |
| Software Name | Purpose |
|---|---|
| SCALEPACK | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data collection |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| MOLREP | phasing |