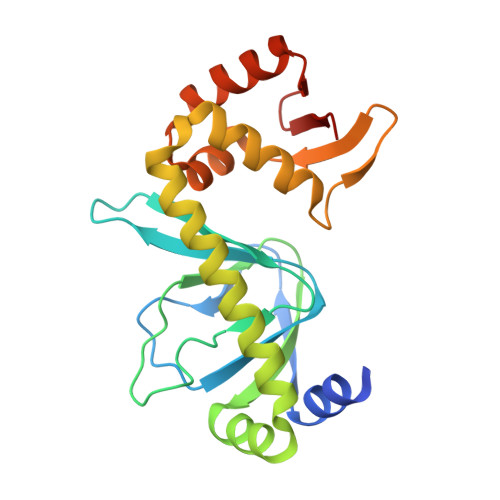
Structure of catabolite activator protein with cobalt(II) and sulfate.
Rao, R.R., Lawson, C.L.(2014) Acta Crystallogr Sect F Struct Biol Cryst Commun 70: 560-563
- PubMed: 24817710 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14005366
- Primary Citation Related Structures:
4FT8 - PubMed Abstract:
The crystal structure of cyclic AMP-catabolite activator protein (CAP) from Escherichia coli containing cobalt(II) chloride and ammonium sulfate is reported at 1.97 Å resolution. Each of the two CAP subunits in the asymmetric unit binds one cobalt(II) ion, in each case coordinated by N-terminal domain residues His19, His21 and Glu96 plus an additional acidic residue contributed via a crystal contact. The three identified N-terminal domain cobalt-binding residues are part of a region of CAP that is important for transcription activation at class II CAP-dependent promoters. Sulfate anions mediate additional crystal lattice contacts and occupy sites corresponding to DNA backbone phosphate positions in CAP-DNA complex structures.
- Department of Chemistry and Chemical Biology, Rutgers University, Piscataway, NJ 08854, USA.
Organizational Affiliation: