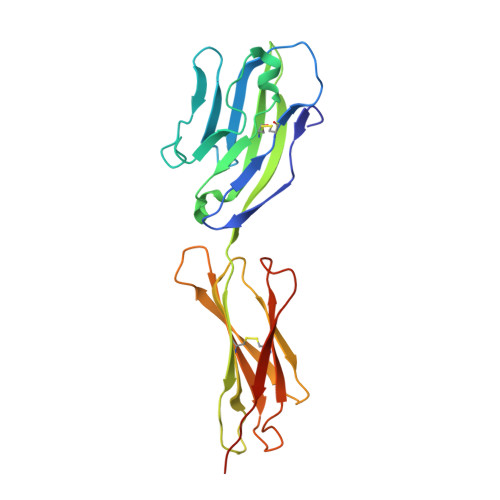
Nectin ectodomain structures reveal a canonical adhesive interface.
Harrison, O.J., Vendome, J., Brasch, J., Jin, X., Hong, S., Katsamba, P.S., Ahlsen, G., Troyanovsky, R.B., Troyanovsky, S.M., Honig, B., Shapiro, L.(2012) Nat Struct Mol Biol 19: 906-915
- PubMed: 22902367 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2366
- Primary Citation Related Structures:
4FMF, 4FMK, 4FN0, 4FOM, 4FQP, 4FRW, 4FS0 - PubMed Abstract:
Nectins are immunoglobulin superfamily glycoproteins that mediate intercellular adhesion in many vertebrate tissues. Homophilic and heterophilic interactions between nectin family members help mediate tissue patterning. We determined the homophilic binding affinities and heterophilic specificities of all four nectins and the related protein nectin-like 5 (Necl-5) from human and mouse, revealing a range of homophilic interaction strengths and a defined heterophilic specificity pattern. To understand the molecular basis of their adhesion and specificity, we determined the crystal structures of natively glycosylated full ectodomains or adhesive fragments of all four nectins and Necl-5. All of the crystal structures revealed dimeric nectins bound through a stereotyped interface that was previously proposed to represent a cis dimer. However, conservation of this interface and the results of targeted cross-linking experiments showed that this dimer probably represents the adhesive trans interaction. The structure of the dimer provides a simple molecular explanation for the adhesive binding specificity of nectins.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, New York, USA.
Organizational Affiliation: