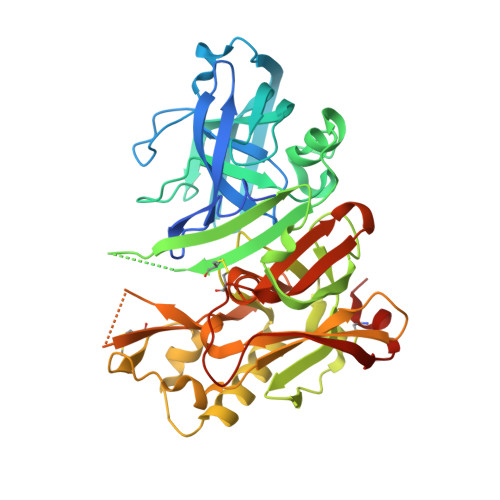
Structure- and Property-Based Design of Aminooxazoline Xanthenes as Selective, Orally Efficacious, and CNS Penetrable BACE Inhibitors for the Treatment of Alzheimer's Disease.
Huang, H., La, D.S., Cheng, A.C., Whittington, D.A., Patel, V.F., Chen, K., Dineen, T.A., Epstein, O., Graceffa, R., Hickman, D., Kiang, Y.H., Louie, S., Luo, Y., Wahl, R.C., Wen, P.H., Wood, S., Fremeau, R.T.(2012) J Med Chem 55: 9156-9169
- PubMed: 22928914 Search on PubMed
- DOI: https://doi.org/10.1021/jm300598e
- Primary Citation Related Structures:
4FRI, 4FRJ, 4FRK - PubMed Abstract:
A structure- and property-based drug design approach was employed to identify aminooxazoline xanthenes as potent and selective human β-secretase inhibitors. These compounds exhibited good isolated enzyme, cell potency, and selectivity against the structurally related aspartyl protease cathepsin D. Our efforts resulted in the identification of a potent, orally bioavailable CNS penetrant compound that exhibited in vivo efficacy. A single oral dose of compound 11a resulted in a significant reduction of CNS Aβ40 in naive rats.
- Department of Medicinal Chemistry, Amgen Inc., 360 Binney Street, Cambridge, Massachusetts 02142, United States. hongbing.huang@amgen.com
Organizational Affiliation: