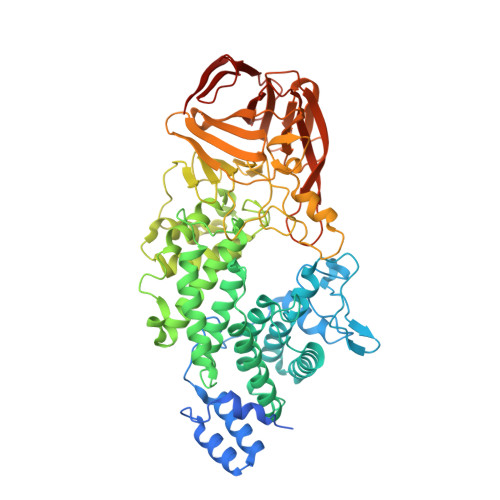
Structural basis of heparan sulfate-specific degradation by heparinase III.
Dong, W., Lu, W., McKeehan, W.L., Luo, Y., Ye, S.(2012) Protein Cell 3: 950-961
- PubMed: 23011846 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s13238-012-2056-z
- Primary Citation Related Structures:
4FNV - PubMed Abstract:
Heparinase III (HepIII) is a 73-kDa polysaccharide lyase (PL) that degrades the heparan sulfate (HS) polysaccharides at sulfate-rare regions, which are important co-factors for a vast array of functional distinct proteins including the well-characterized antithrombin and the FGF/FGFR signal transduction system. It functions in cleaving metazoan heparan sulfate (HS) and providing carbon, nitrogen and sulfate sources for host microorganisms. It has long been used to deduce the structure of HS and heparin motifs; however, the structure of its own is unknown. Here we report the crystal structure of the HepIII from Bacteroides thetaiotaomicron at a resolution of 1.6 Å. The overall architecture of HepIII belongs to the (α/α)₅ toroid subclass with an N-terminal toroid-like domain and a C-terminal β-sandwich domain. Analysis of this high-resolution structure allows us to identify a potential HS substrate binding site in a tunnel between the two domains. A tetrasaccharide substrate bound model suggests an elimination mechanism in the HS degradation. Asn260 and His464 neutralize the carboxylic group, whereas Tyr314 serves both as a general base in C-5 proton abstraction, and a general acid in a proton donation to reconstitute the terminal hydroxyl group, respectively. The structure of HepIII and the proposed reaction model provide a molecular basis for its potential practical utilization and the mechanism of its eliminative degradation for HS polysaccarides.
- Life Sciences Institute, Zhejiang University, Hangzhou 310058, China.
Organizational Affiliation: