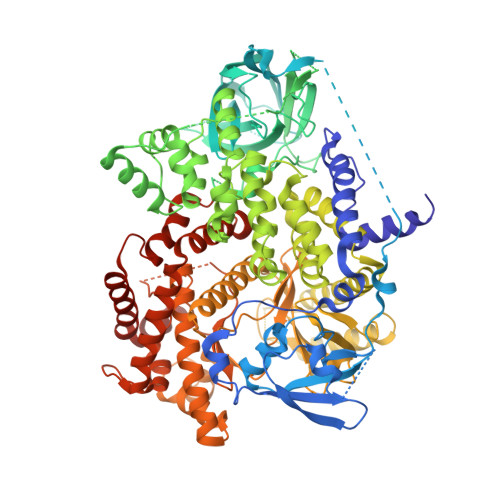
Discovery and in Vivo Evaluation of Dual PI3K-beta/delta inhibitors
Gonzalez-Lopez de Turiso, F., Shin, Y., Brown, M., Cardozo, M., Chen, Y., Fong, D., Hao, X., He, X., Henne, K., Hu, Y.L., Johnson, M.G., Kohn, T., Lohman, J., McBride, H.J., McGee, L.R., Medina, J.C., Metz, D., Miner, K., Mohn, D., Pattaropong, V., Seganish, J., Simard, J.L., Wannberg, S., Whittington, D.A., Yu, G., Cushing, T.D.(2012) J Med Chem 55: 7667-7685
- PubMed: 22876881 Search on PubMed
- DOI: https://doi.org/10.1021/jm300679u
- Primary Citation Related Structures:
4FJY, 4FJZ - PubMed Abstract:
Structure-based rational design led to the synthesis of a novel series of potent PI3K inhibitors. The optimized pyrrolopyridine analogue 63 was a potent and selective PI3Kβ/δ dual inhibitor that displayed suitable physicochemical properties and pharmacokinetic profile for animal studies. Analogue 63 was found to be efficacious in animal models of inflammation including a keyhole limpet hemocyanin (KLH) study and a collagen-induced arthritis (CIA) disease model of rheumatoid arthritis. These studies highlight the potential therapeutic value of inhibiting both the PI3Kβ and δ isoforms in the treatment of a number of inflammatory diseases.
- Department of Therapeutic Discovery, Amgen Inc., 1120 Veterans Boulevard, South San Francisco, California 94080, USA. felgonza@amgen.com
Organizational Affiliation: