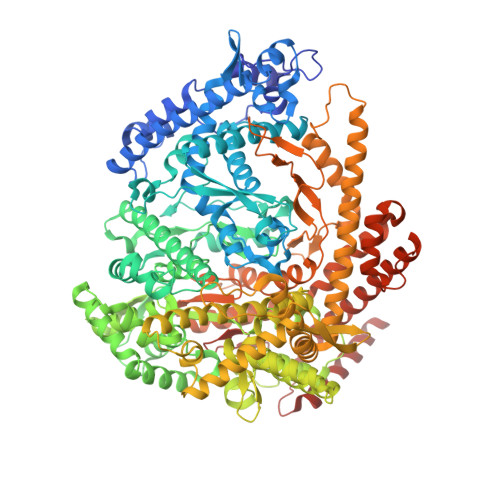
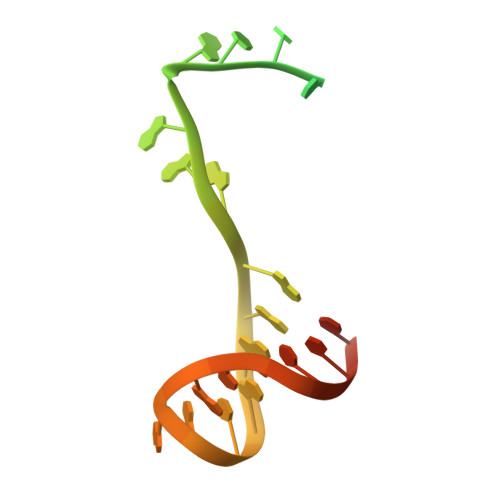
Watching the Bacteriophage N4 RNA Polymerase Transcription by Time-dependent Soak-trigger-freeze X-ray Crystallography.
Basu, R.S., Murakami, K.S.(2013) J Biological Chem 288: 3305-3311
- PubMed: 23235152 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.387712
- Primary Citation Related Structures:
4FF1, 4FF2, 4FF3, 4FF4 - PubMed Abstract:
The challenge for structural biology is to understand atomic-level macromolecular motions during enzymatic reaction. X-ray crystallography can reveal high resolution structures; however, one perceived limitation is that it reveals only static views. Here we use time-dependent soak-trigger-freeze X-ray crystallography, namely, soaking nucleotide and divalent metal into the bacteriophage RNA polymerase (RNAP)-promoter DNA complex crystals to trigger the nucleotidyl transfer reaction and freezing crystals at different time points, to capture real-time intermediates in the pathway of transcription. In each crystal structure, different intensities and shapes of electron density maps corresponding to the nucleotide and metal were revealed at the RNAP active site which allow watching the nucleotide and metal bindings and the phosphodiester bond formation in a time perspective. Our study provides the temporal order of substrate assembly and metal co-factor binding at the active site of enzyme which completes our understanding of the two-metal-ion mechanism and fidelity mechanism in single-subunit RNAPs. The nucleotide-binding metal (Me(B)) is coordinated at the active site prior to the catalytic metal (Me(A)). Me(A) coordination is only temporal, established just before and dissociated immediately after phosphodiester bond formation. We captured these elusive intermediates exploiting the slow enzymatic reaction in crystallo. These results demonstrate that the simple time-dependent soak-trigger-freeze X-ray crystallography offers a direct means for monitoring enzymatic reactions.
- Department of Biochemistry and Molecular Biology, The Center for RNA Molecular Biology, The Pennsylvania State University, University Park, Pennsylvania 16802, USA.
Organizational Affiliation: