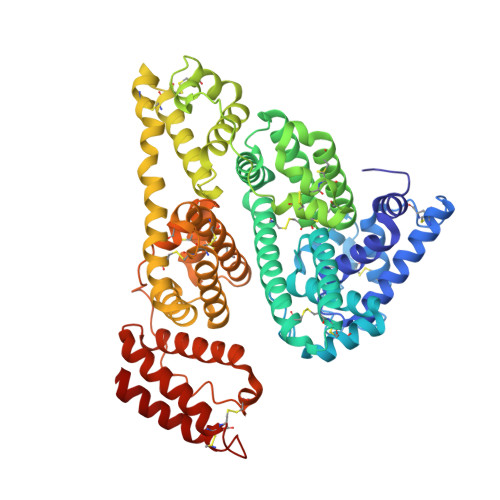
Structures of bovine, equine and leporine serum albumin.
Bujacz, A.(2012) Acta Crystallogr D Biol Crystallogr 68: 1278-1289
- PubMed: 22993082 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912027047
- Primary Citation Related Structures:
4F5S, 4F5T, 4F5U, 4F5V - PubMed Abstract:
Serum albumin first appeared in early vertebrates and is present in the plasma of all mammals. Its canonical structure supported by a conserved set of disulfide bridges is maintained in all mammalian serum albumins and any changes in sequence are highly correlated with evolution of the species. Previous structural investigations of mammalian serum albumins have only concentrated on human serum albumin (HSA), most likely as a consequence of crystallization and diffraction difficulties. Here, the crystal structures of serum albumins isolated from bovine, equine and leporine blood plasma are reported. The structure of bovine serum albumin (BSA) was determined at 2.47 Å resolution, two crystal structures of equine serum albumin (ESA) were determined at resolutions of 2.32 and 2.04 Å, and that of leporine serum albumin (LSA) was determined at 2.27 Å resolution. These structures were compared in detail with the structure of HSA. The ligand-binding pockets in BSA, ESA and LSA revealed different amino-acid compositions and conformations in comparison to HSA in some cases; however, much more significant differences were observed on the surface of the molecules. BSA, which is one of the most extensively utilized proteins in laboratory practice and is used as an HSA substitute in many experiments, exhibits only 75.8% identity compared with HSA. The higher resolution crystal structure of ESA highlights the binding properties of this protein because it includes several bound compounds from the crystallization solution that provide additional structural information about potential ligand-binding pockets.
- Institute of Technical Biochemistry, Lodz University of Technology, Stefanowskiego 4/10, 90-924 Lodz, Poland.
Organizational Affiliation: