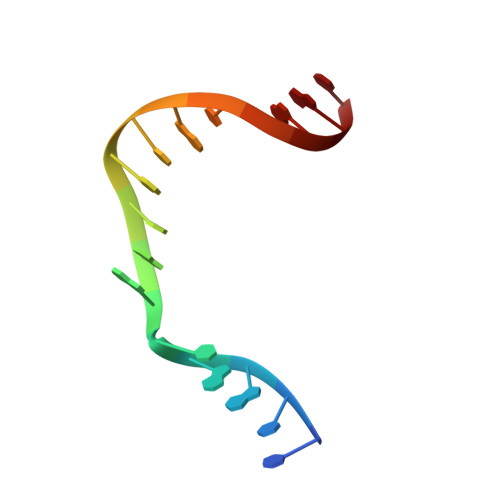
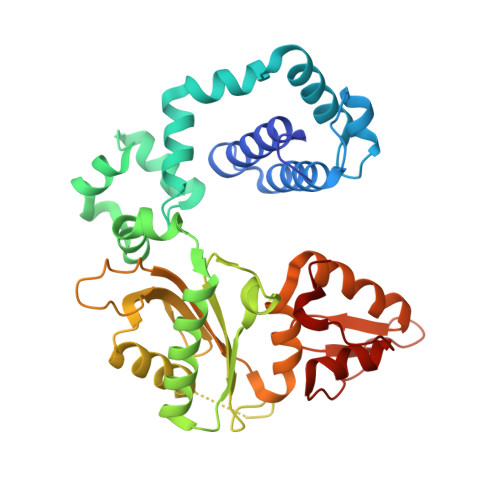
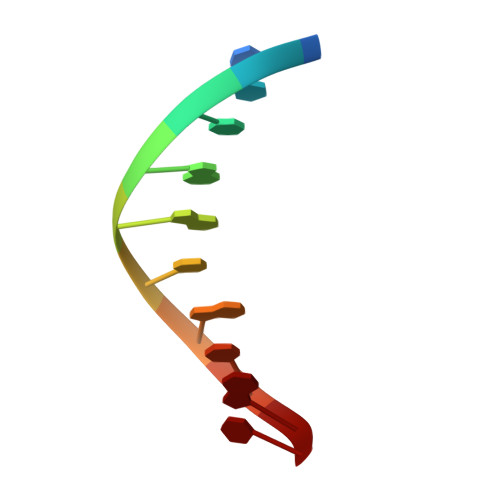
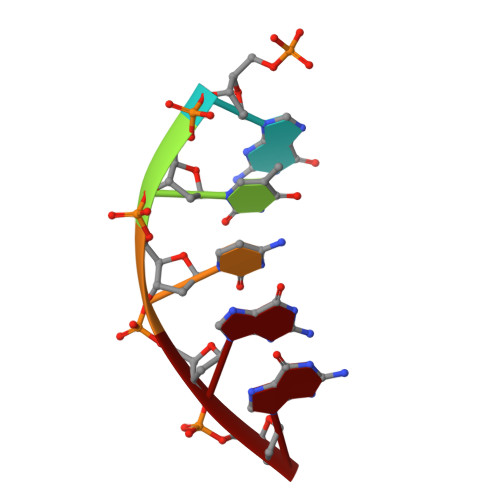
Structures of dNTP Intermediate States during DNA Polymerase Active Site Assembly.
Freudenthal, B.D., Beard, W.A., Wilson, S.H.(2012) Structure 20: 1829-1837
- PubMed: 22959623 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2012.08.008
- Primary Citation Related Structures:
4F5N, 4F5O, 4F5P, 4F5Q, 4F5R - PubMed Abstract:
DNA polymerase and substrate conformational changes are essential for high-fidelity DNA synthesis. Structures of DNA polymerase (pol) β in complex with DNA show the enzyme in an "open" conformation. Subsequent to binding the nucleotide, the polymerase "closes" around the nascent base pair with two metals positioned for chemistry. However, structures of substrate/active site intermediates prior to closure are lacking. By destabilizing the closed complex, we determined unique ternary complex structures of pol β with correct and incorrect incoming nucleotides bound to the open conformation. These structures reveal that Watson-Crick hydrogen bonding is assessed upon initial complex formation. Importantly, nucleotide-bound states representing intermediate metal coordination states occur with active site assembly. The correct, but not incorrect, nucleotide maintains Watson-Crick hydrogen bonds during interconversion of these states. These structures indicate that the triphosphate of the incoming nucleotide undergoes rearrangement prior to closure, providing an opportunity to deter misinsertion and increase fidelity.
- Laboratory of Structural Biology, National Institute of Environmental Health Sciences, National Institutes of Health, P.O. Box 12233, Research Triangle Park, NC 27709-2233, USA.
Organizational Affiliation: