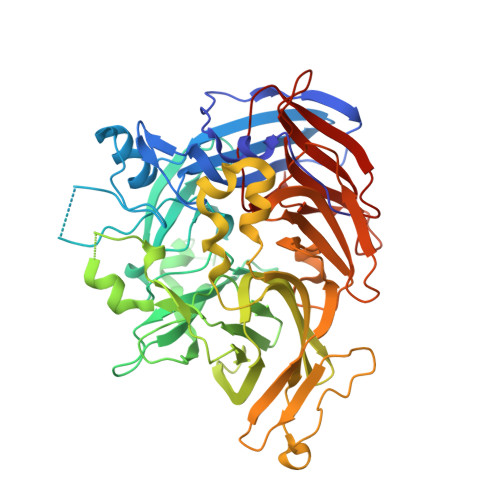
Structure of RPE65 isomerase in a lipidic matrix reveals roles for phospholipids and iron in catalysis.
Kiser, P.D., Farquhar, E.R., Shi, W., Sui, X., Chance, M.R., Palczewski, K.(2012) Proc Natl Acad Sci U S A 109: E2747-E2756
- PubMed: 23012475 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1212025109
- Primary Citation Related Structures:
4F2Z, 4F30, 4F3A, 4F3D - PubMed Abstract:
RPE65 is a key metalloenzyme responsible for maintaining visual function in vertebrates. Despite extensive research on this membrane-bound retinoid isomerase, fundamental questions regarding its enzymology remain unanswered. Here, we report the crystal structure of RPE65 in a membrane-like environment. These crystals, obtained from enzymatically active, nondelipidated protein, displayed an unusual packing arrangement wherein RPE65 is embedded in a lipid-detergent sheet. Structural differences between delipidated and nondelipidated RPE65 uncovered key residues involved in substrate uptake and processing. Complementary iron K-edge X-ray absorption spectroscopy data established that RPE65 as isolated contained a divalent iron center and demonstrated the presence of a tightly bound ligand consistent with a coordinated carboxylate group. These results support the hypothesis that the Lewis acidity of iron could be used to promote ester dissociation and generation of a carbocation intermediate required for retinoid isomerization.
- Department of Pharmacology, School of Medicine, Case Western Reserve University, Cleveland, OH 44106, USA. pdk7@case.edu
Organizational Affiliation: