Synthesis and Antisense Properties of Fluoro Cyclohexenyl Nucleic Acid (F-CeNA), a Nuclease Stable Mimic of 2'-Fluoro RNA.
Seth, P.P., Yu, J., Jazayeri, A., Pallan, P.S., Allerson, C.R., Ostergaard, M.E., Liu, F., Herdewijn, P., Egli, M., Swayze, E.E.(2012) J Org Chem 77: 5074-5085
- PubMed: 22591005 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jo300594b
- Primary Citation Related Structures:
4F2X, 4F2Y - PubMed Abstract:
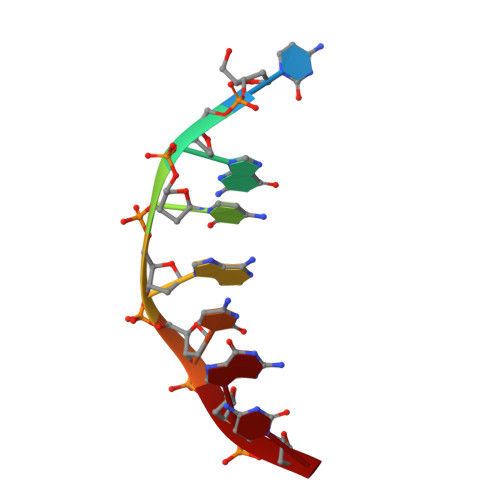
We report the design and synthesis of 2'-fluoro cyclohexenyl nucleic acid (F-CeNA) pyrimidine phosphoramidites and the synthesis and biophysical, structural, and biological evaluation of modified oligonucleotides. The synthesis of the nucleoside phosphoramidites was accomplished in multigram quantities starting from commercially available methyl-D-mannose pyranoside. Installation of the fluorine atom was accomplished using nonafluorobutanesulfonyl fluoride, and the cyclohexenyl ring system was assembled by means of a palladium-catalyzed Ferrier rearrangement. Installation of the nucleobase was carried out under Mitsunobu conditions followed by standard protecting group manipulations to provide the desired pyrimidine phosphoramidites. Biophysical evaluation indicated that F-CeNA shows behavior similar to that of a 2'-modified nucleotide, and duplexes with RNA showed slightly lower duplex thermostability as compared to that of the more rigid 3'-fluoro hexitol nucleic acid (FHNA). However, F-CeNA modified oligonucleotides were significantly more stable against digestion by snake venom phosphodiesterases (SVPD) as compared to unmodified DNA, 2'-fluoro RNA (FRNA), 2'-methoxyethyl RNA (MOE), and FHNA modified oligonucleotides. Examination of crystal structures of a modified DNA heptamer duplex d(GCG)-T*-d(GCG):d(CGCACGC) by X-ray crystallography indicated that the cyclohexenyl ring system exhibits both the (3)H(2) and (2)H(3) conformations, similar to the C3'-endo/C2'-endo conformation equilibrium seen in natural furanose nucleosides. In the (2)H(3) conformation, the equatorial fluorine engages in a relatively close contact with C8 (2.94 Å) of the 3'-adjacent dG nucleotide that may represent a pseudo hydrogen bond. In contrast, the cyclohexenyl ring of F-CeNA was found to exist exclusively in the (3)H(2) (C3'-endo like) conformation in the crystal structure of the modified A-form DNA decamer duplex [d(GCGTA)-T*-d(ACGC)](2.) In an animal experiment, a 16-mer F-CeNA gapmer ASO showed similar RNA affinity but significantly improved activity compared to that of a sequence matched MOE ASO, thus establishing F-CeNA as a useful modification for antisense applications.
- Isis Pharmaceuticals, 2855 Gazelle Court, Carlsbad, California 92010, United States. pseth@isisph.com
Organizational Affiliation: