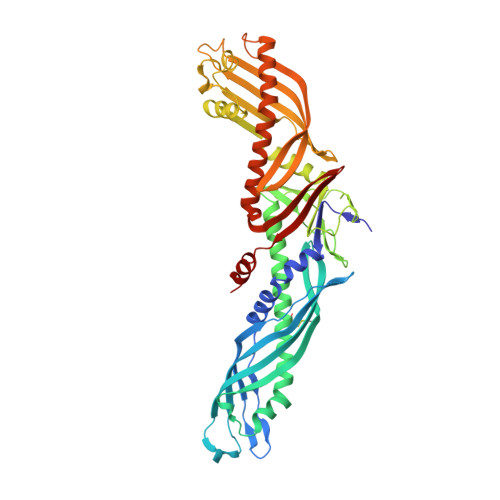
Crystal structures of cholesteryl ester transfer protein in complex with inhibitors.
Liu, S., Mistry, A., Reynolds, J.M., Lloyd, D.B., Griffor, M.C., Perry, D.A., Ruggeri, R.B., Clark, R.W., Qiu, X.(2012) J Biological Chem 287: 37321-37329
- PubMed: 22961980 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.380063
- Primary Citation Related Structures:
4EWS, 4F2A - PubMed Abstract:
Human plasma cholesteryl ester transfer protein (CETP) transports cholesteryl ester from the antiatherogenic high-density lipoproteins (HDL) to the proatherogenic low-density and very low-density lipoproteins (LDL and VLDL). Inhibition of CETP has been shown to raise human plasma HDL cholesterol (HDL-C) levels and is potentially a novel approach for the prevention of cardiovascular diseases. Here, we report the crystal structures of CETP in complex with torcetrapib, a CETP inhibitor that has been tested in phase 3 clinical trials, and compound 2, an analog from a structurally distinct inhibitor series. In both crystal structures, the inhibitors are buried deeply within the protein, shifting the bound cholesteryl ester in the N-terminal pocket of the long hydrophobic tunnel and displacing the phospholipid from that pocket. The lipids in the C-terminal pocket of the hydrophobic tunnel remain unchanged. The inhibitors are positioned near the narrowing neck of the hydrophobic tunnel of CETP and thus block the connection between the N- and C-terminal pockets. These structures illuminate the unusual inhibition mechanism of these compounds and support the tunnel mechanism for neutral lipid transfer by CETP. These highly lipophilic inhibitors bind mainly through extensive hydrophobic interactions with the protein and the shifted cholesteryl ester molecule. However, polar residues, such as Ser-230 and His-232, are also found in the inhibitor binding site. An enhanced understanding of the inhibitor binding site may provide opportunities to design novel CETP inhibitors possessing more drug-like physical properties, distinct modes of action, or alternative pharmacological profiles.
- Department of Structural Biology & Biophysics, Pfizer Groton Laboratories, Groton, Connecticut 06340, USA. shenping.liu@pfizer.com
Organizational Affiliation: